Electron microscopy: from the dark ages to a bright future
Posted by Lucy Collinson, on 21 July 2020
Good sample preparation is, as every microscopist knows, the key to delivering sound results from an imaging experiment. In the digital age, and with the advent of big data, image analysis is also critical to extract meaningful quantitative results from image data. Indeed nowadays, the microscope itself is often the most well-developed and user-friendly part of an imaging workflow, fine-tuned over decades of academic and industrial investment and innovation.
Electron microscopy (EM) is a specific case in point. Learning to use a benchtop scanning electron microscope is now almost as easy as learning to use an app on a tablet. However, sample preparation is inherently complex because of the competing requirements to image the sample in a vacuum and to image it as close to its original living state as possible. Likewise, image analysis in EM is particularly challenging because one of the greatest benefits of the electron microscope – the ability to capture many different classes of biological structure in a heterogenous sample in a single image – is at odds with current (human and software) capabilities to recognise and classify all structures in that information-dense image.
Historically, EM has been a more ‘hidden’ branch of imaging than light microscopy in the life sciences, partly because of the challenges of sample preparation (toxic chemicals, numerous steps, long timeframes and miniscule samples), but perhaps also because of the complexity of using early microscopes and the physical requirements for hosting them. They are usually large, heavy, and require a stable environment that is low in physical and acoustic vibrations, electromagnetic fields and ambient light. These environments tend to be found in the lowest basement levels of buildings, which contributed to the relative isolation of the EM community in the late 20th Century.
From the beginning of the 21st Century, EM experienced a revival, leading to two distinct revolutions in high-resolution imaging of macromolecules and virus particles (cryo EM), as well as in three dimensional characterisation of cells and tissues (volume EM). The cryo EM and volume EM communities managed to break out of their (physical and metaphorical) isolated basements, and recruited a new generation of visible, collaborative and well-networked imaging scientists, quickly delivering huge advances in our understanding of biological systems at different scales. In the process, the field has seen an explosion of new technology, including (i) the democratisation of transmission EMs (TEMs) and scanning EMs (SEMs) through advances in user-friendly software and digital detectors, (ii) an increased portability and accessibility via minituarisation to create benchtop SEMs and TEMs, (iii) automation of 200 keV and 300 kV cryo TEMs for single-particle and tomography experiments, (iv) automation of serial section imaging in TEMs via software and workflow advances, as well as (v) an array of tomography and blockface imaging methods (serial block face SEM (SBF SEM) and focused ion beam (FIB SEM)) for volume EM of resin-embedded cells and tissues. Even as I write, new higher-throughput electron-imaging technologies are becoming available, including multibeam SEMs, gridTape TEM and plasma FIBs.
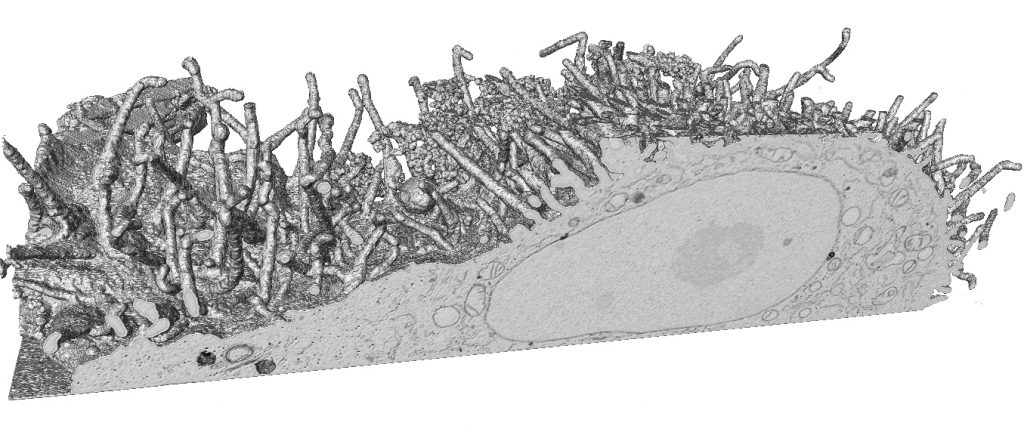
Back in the late 20th Century, EM had fallen out of fashion, largely due to the revolution in confocal light-microscopy that went hand-in-hand with the rise of green fluorescent protein as a tool for spatial molecular biology. EM only recovered its status as the go-to method for structural cell biology when the limitations of fluorescence microscopy became clear. Later came a series of inspired breakthroughs that rendered the Abbe limit of maximum resolution meaningless and delivered a new era of super resolution light microscopy, but by then, EM was well established and its unique ability to image the complex structural interplay of all cellular components (and not just those tagged with a fluorescent protein) was clear. In this respect, perhaps the greatest challenge to EM comes from the burgeoning developments and application of X-ray microscopy (XRM) in the life sciences. XRM is capable of revealing cell and tissue structure with a remarkably similar ‘look-and-feel’ to images generated by EM. Having already had huge impact in the area of macromolecular crystallography, synchrotron radiation is now being used to image cells and tissues on soft and hard X-ray tomography beamlines, and XRM has reached smaller institutional imaging facilities via lab-scale micro-computed tomography (microCT) where it is used for nano-anatomy and non-destructive targeting of regions of interest in tissues for subsequent EM image acquisition.
But to separate light, electron and X-ray microscopes and their respective communities, and to pitch them against each other in a ‘battle of the imaging modalities’ is to miss the point. The future of bioimaging is in the crosstalk between modalities, in the murky and hard-to-navigate waters of the interdisciplinary divide. While ‘omics’ communities attempt to reduce complexity through advances in single-cell technologies, imaging communities are attempting to broaden their existing single-cell technologies to tackle the complex spatial systems biology of entire organs like the brain, and pathological tissues such as tumours. Given the age-old compromise between sample size and resolution, and between imaging structure and function, these challenges will only be met by designing new multimodal and correlative workflows that ‘daisy-chain’ microscopes together to capture images of nanoscale biological processes across scales within a single organism, be that an experimental model organism or the human patient.
With new studies reporting a broader imaging and spatial ‘omics’ data integration to understand the diversity of individuals in a species, we are getting closer to an achievable practical systems-biology approach, in which imaging and EM play a fundamental role. To loop us back to the beginning of this piece, the missing links in these multimodal workflows lie firmly in the domains of sample preparation and data analysis, and this is where our community efforts must focus next, through collaboration, data sharing and open science.

Lucy Collinson
Director of EM-STP
The Francis Crick Institute,
1 Midland Road, London, NW1 1AT, UK


 (6 votes, average: 1.00 out of 1)
(6 votes, average: 1.00 out of 1)
Great article but I am always surprised and disappointed that Energy Filtering Transmission Electron Microscopy (EFTEM)/Electron Spectroscopic Imaging (ESI) never gets discussed. Work by David Bazett-Jones and now Michael Hendzel should be included in this. The use of this technique for chromatin/nuclear imaging is highly under utilized considering how many high resolution microscopes are now equipped with energy filters for zero-loss imaging.
Hi Chris, thanks for your comment and apologies for the omission. I agree that spatial elemental and chemical analysis in TEM as well as SEM, nanoSIMS, orbiSIMS etc is a very exciting area, especially with recent improvements in detectors. A good topic for a FocalPlane blog!
great article!! Integrating omics data with EM is a great challenge but fundamental.