Microscopy preprints – New tools and techniques
Posted by FocalPlane, on 14 February 2022
Here is a curated selection of preprints published recently. In this post, we focus specifically on new imaging tools only.
3D multi-color far-red single-molecule localization microscopy with probability-based fluorophore classification. Marijn Siemons, Daphne Jurriens, Carlas Smith, Lukas Kapitein
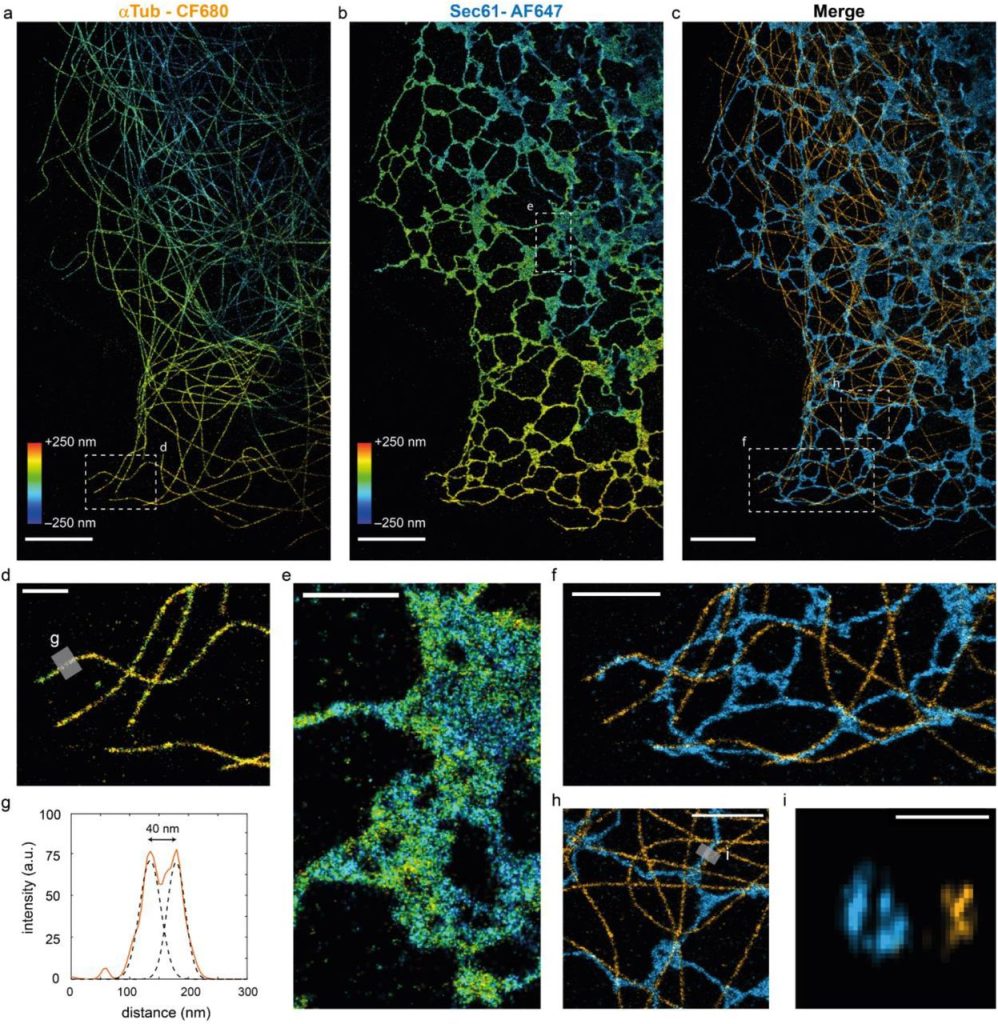
A DNA-based optical force sensor for live-cell applications. Christina Jayachandran, Arindam Ghosh, Meenakshi Prabhune, Jonathan Bath, Andrew J. Turberfield, Lara Hauke, Jörg Enderlein, Florian Rehfeldt, Christoph F. Schmidt
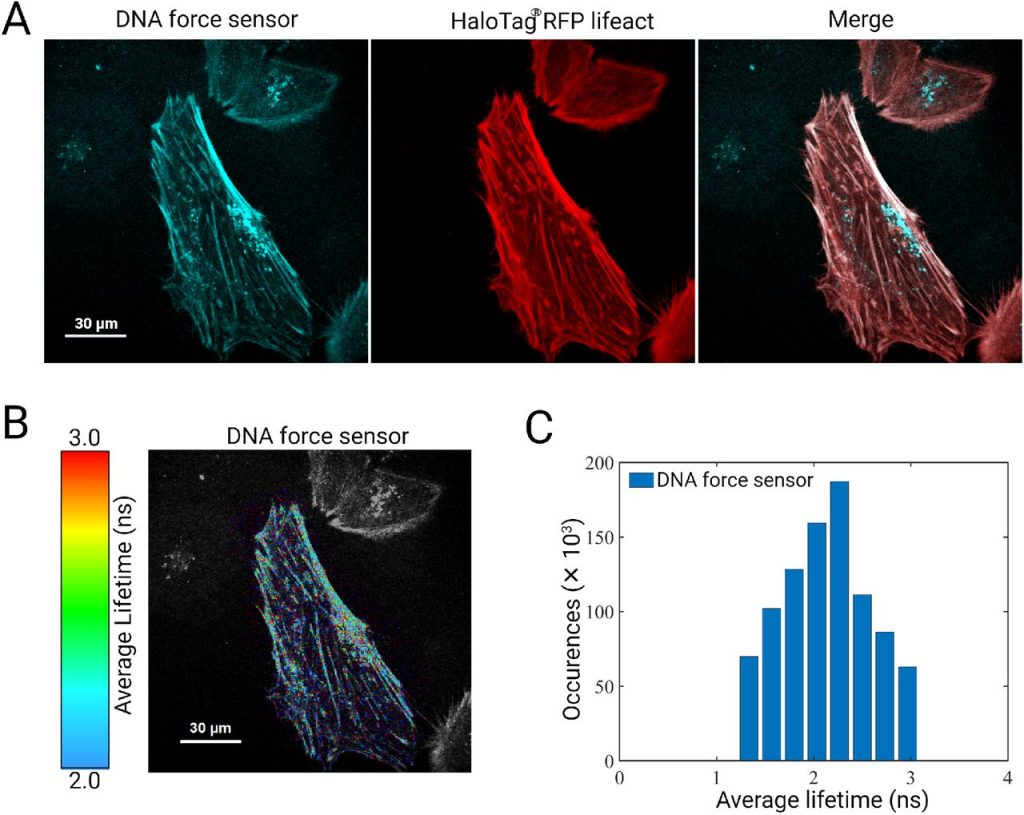
STReTCh: a strategy for facile detection of mechanical forces across proteins in cells. Brian L Zhong, Vipul T Vachharajani, Alexander R Dunn
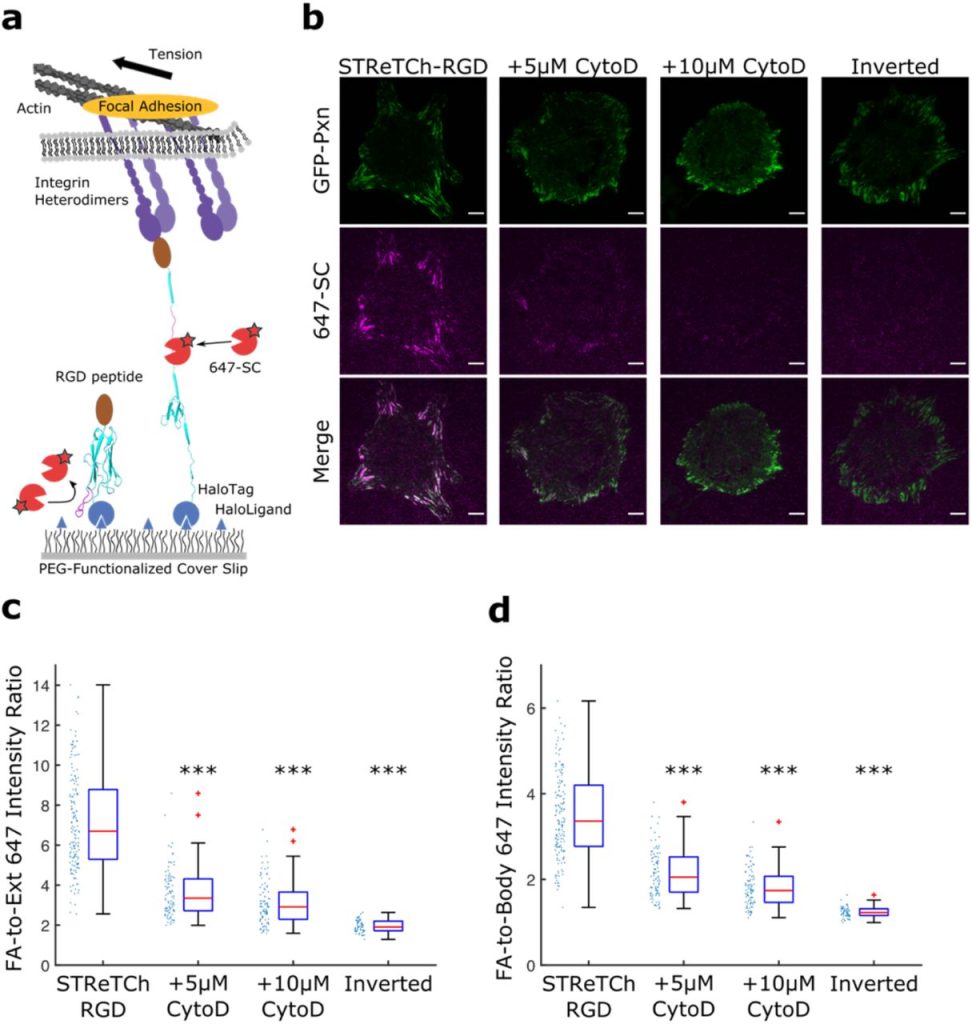
Scalable in situ single-cell profiling by electrophoretic capture of mRNA. Lars Borm, Alejandro Mossi Albiach, Camiel CA Mannens, Jokubas Janusauskas, Ceren Özgün, David Fernández García, Rebecca D Hodge, Ed Lein, Simone Codeluppi, Sten Linnarsson
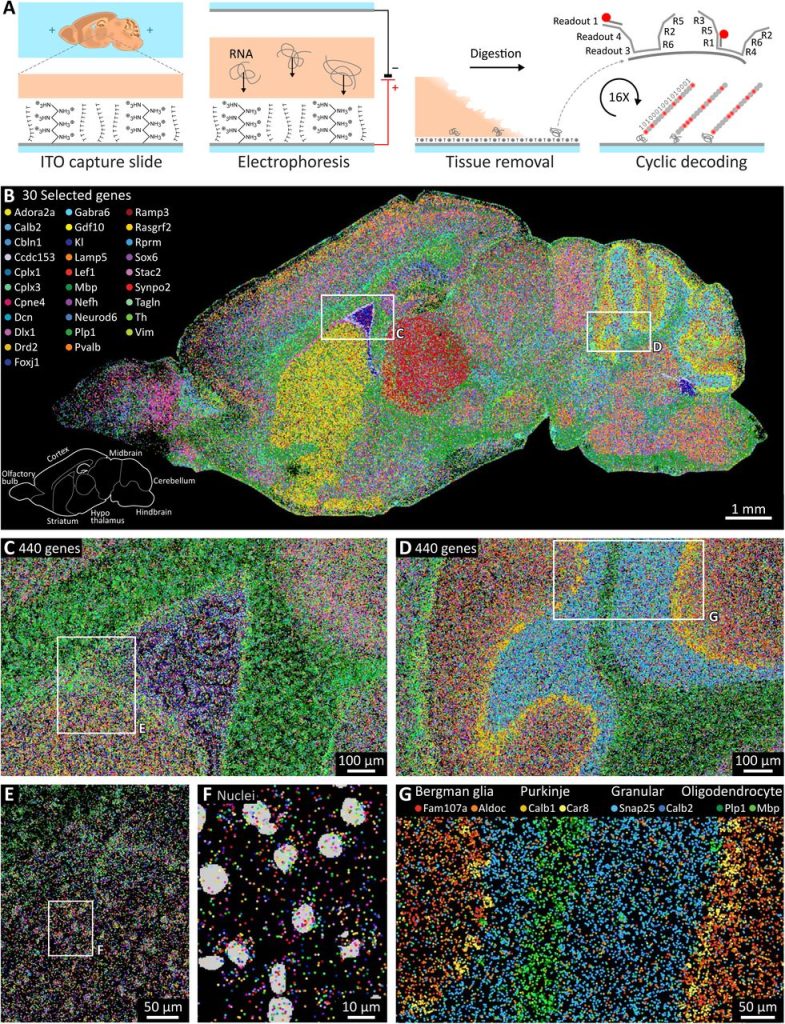
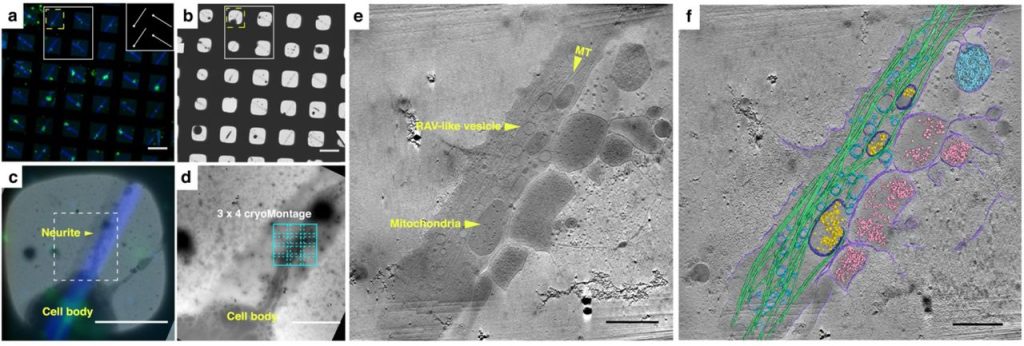
Correlative cryogenic montage electron tomography for comprehensive in-situ whole-cell structural studies. Jie E. Yang, Matthew R. Larson, Bryan S. Sibert, Joseph Y. Kim, Daniel Parrell, Juan C. Sanchez, Victoria Pappas, Anil Kumar, Kai Cai, Keith Thompson, Elizabeth R. Wright

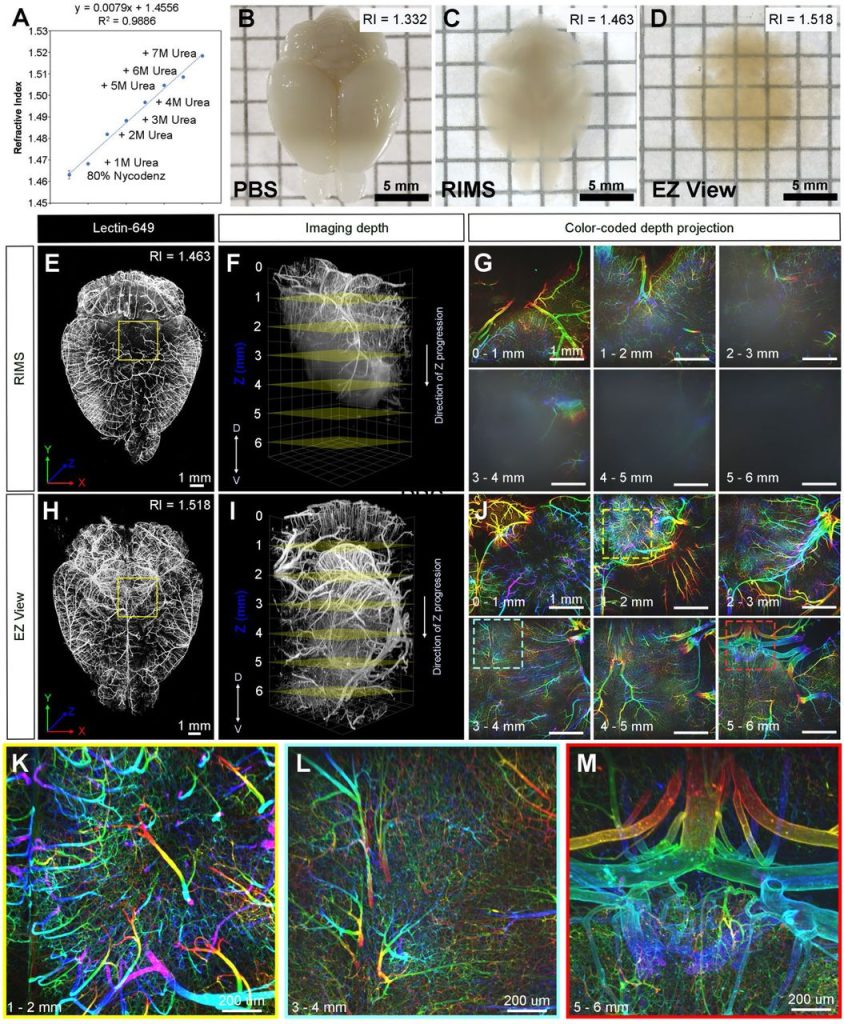
Live-cell fluorescence lifetime multiplexing using synthetic fluorescent probes. Michelle S. Frei, Birgit Koch, Julien Hiblot, Kai Johnsson
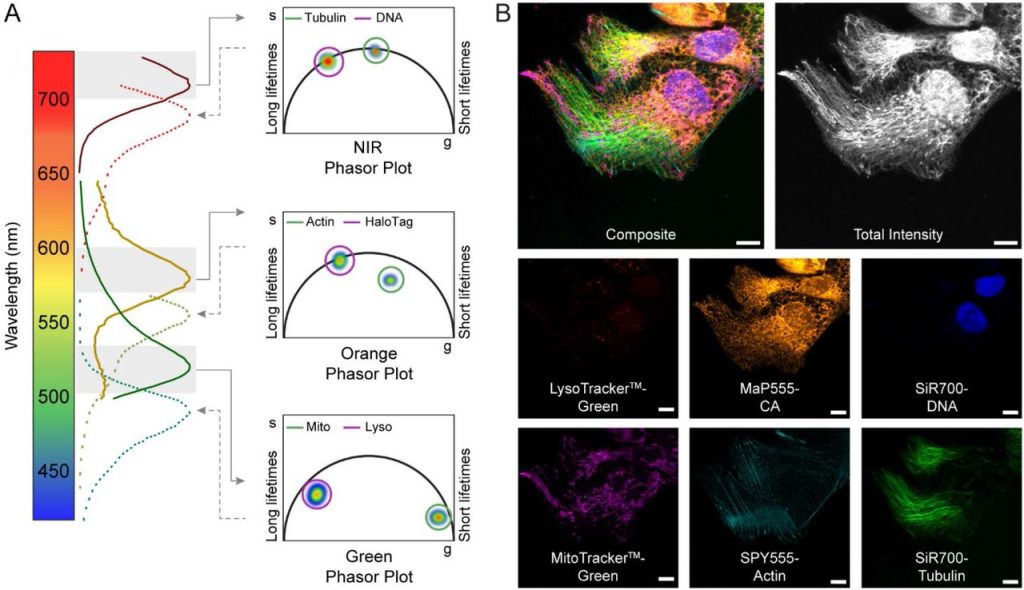
Long-term two-photon imaging of spinal cord in freely behaving mice. Furong Ju, Wenling Jian, Yaning Han, Tianwen Huang, Jin Ke, Zhiheng Liu, Shengyuan Cai, Nan Liu, Liping Wang, Pengfei Wei
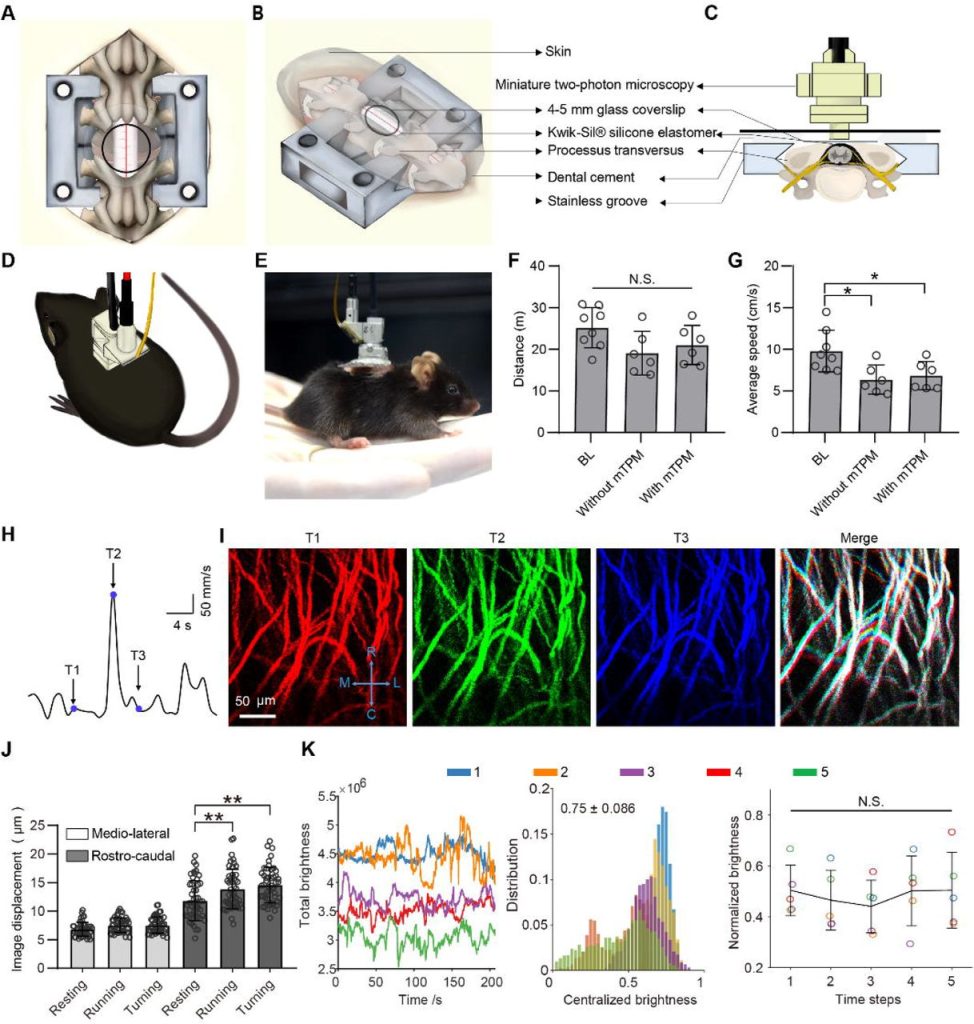
Visualizing Synaptic Dopamine Efflux with a 2D Nanofilm. Chandima Bulumulla, Andrew T Krasley, Deepika Walpita, Abraham G Beyene
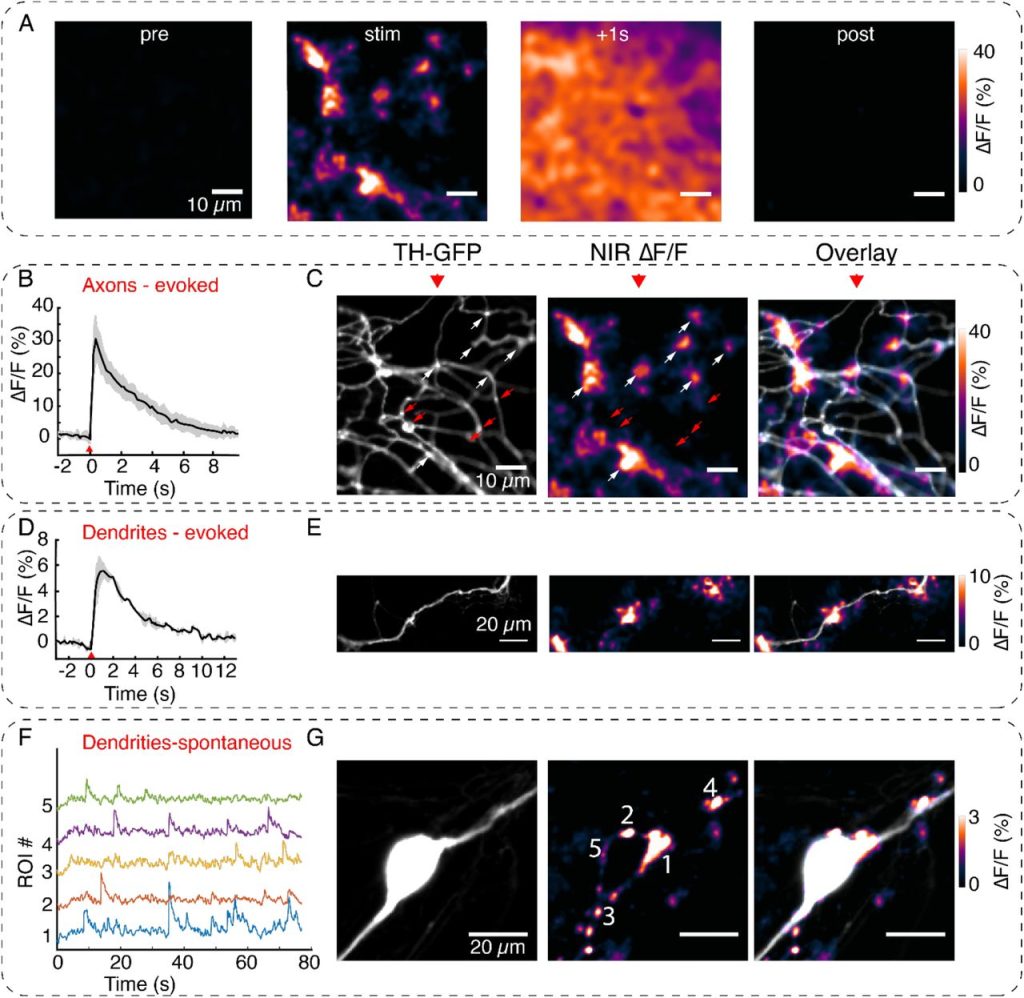
EZ Clear for simple, rapid, and robust mouse whole organ clearing. Chih-Wei Hsu, Juan Cerda III, Jason M Kirk, Williamson D. Turner, Tara L. Rasmussen, Mary E Dickinson, Carlos P. Flores Suarez, Joshua Wythe
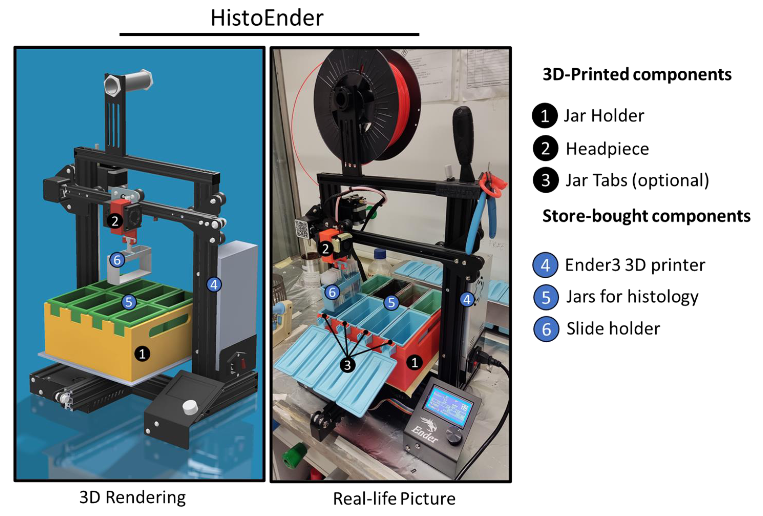
HistoEnder: a 3D printer-based histological slide autostainer that retains 3D printer functions. Marco Ponzetti, Ganga Chinna Rao Devarapu, Nadia Rucc, Armando Carlone, Vittorio Saggiomo
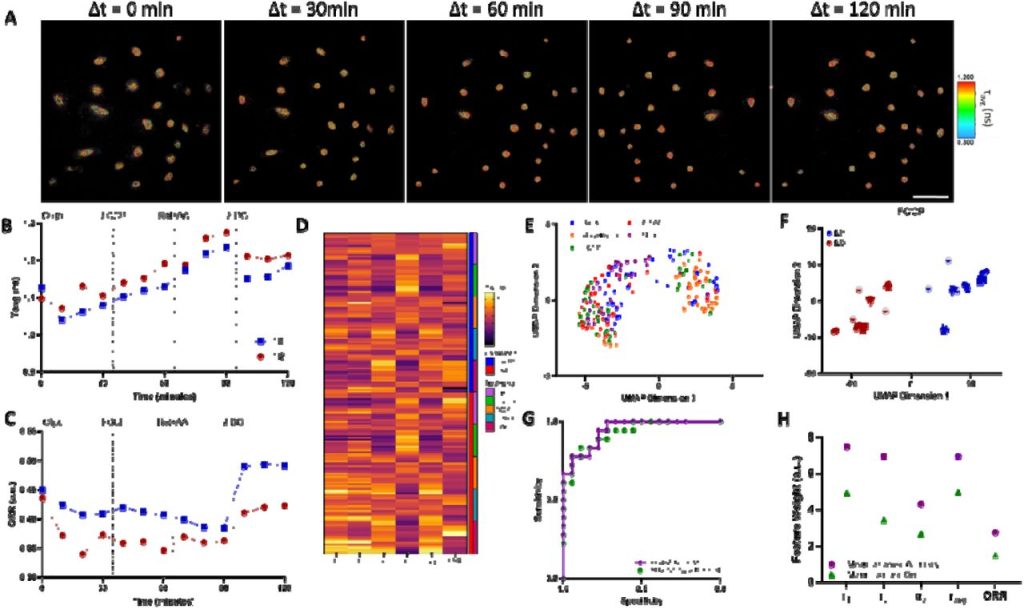
Non-Invasive classification of macrophage polarisation by 2P-FLIM and machine learning. Nuno G.B Neto, Sinead A O’Rourke, Mimi Zhang, Hannah K Fitzgerald, Aisling Dunne, Michael G Monaghan
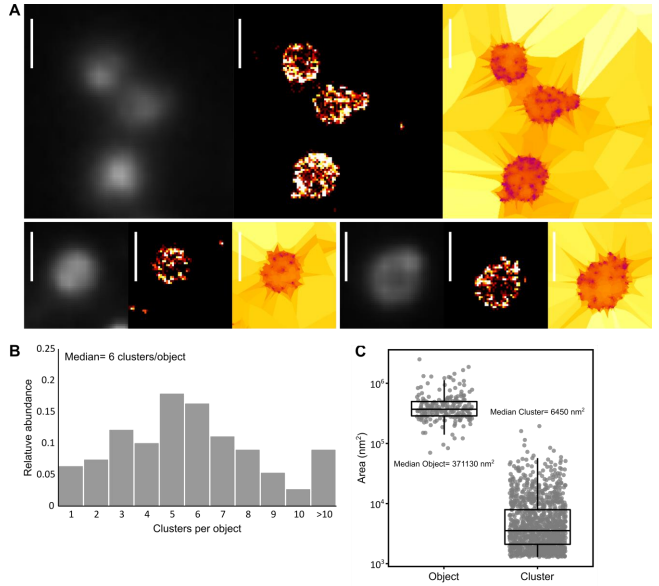
Imaging minimal bacteria at the nanoscale: a reliable and versatile process to perform Single Molecule Localization Microscopy in mycoplasmas. Fabien Rideau, Audrey Villa, Pauline Belzanne, Emeline Verdier, Eric Hosy, Yonathan Arfi
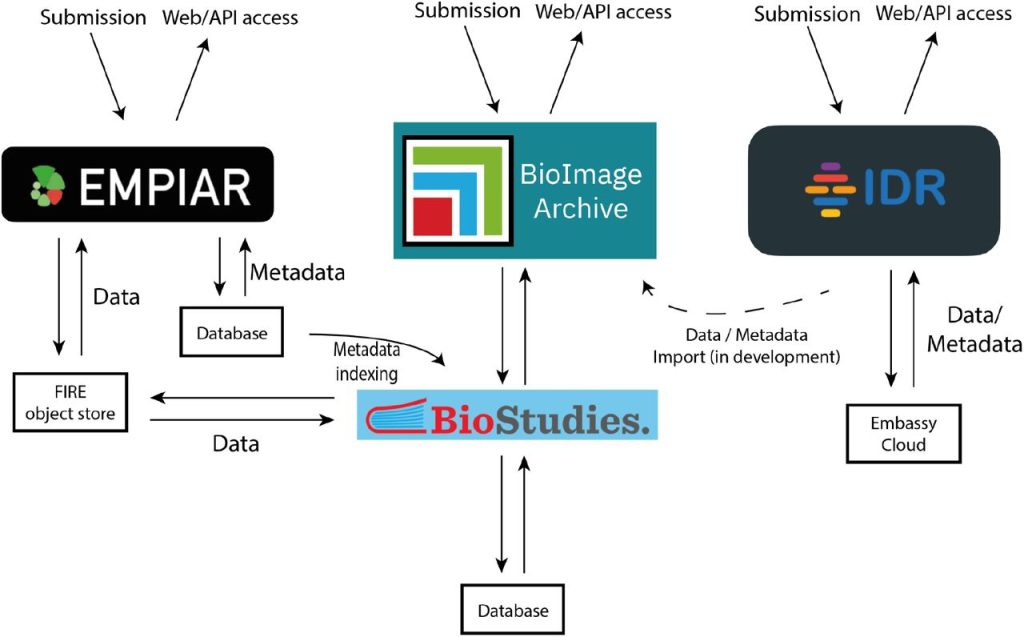
The BioImage Archive – home of life-sciences microscopy data. Matthew Hartley, Gerard Kleywegt, Ardan Patwardhan, Ugis Sarkans, Jason R. Swedlow, Alvis Brazma


 (No Ratings Yet)
(No Ratings Yet)