Affimers – small probes for big imaging experiments
Posted by Laura Hankins, on 9 August 2022
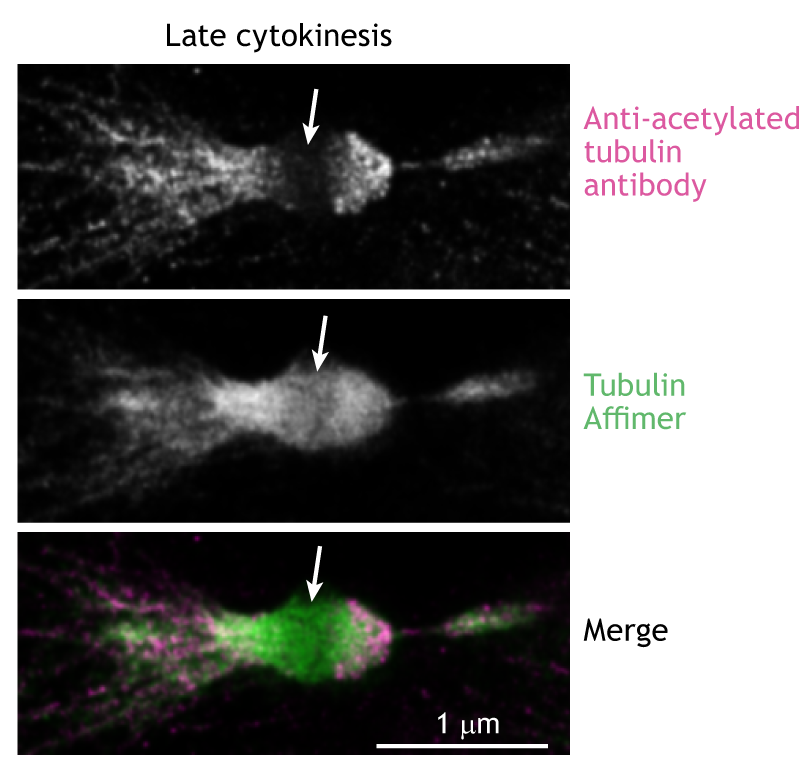
Antibodies have been used to label proteins of interest since the 1970s. However, their large size (around 15 nm long for IgG) makes the penetration of dense structures within the cell a challenge and places the fluorophore away from the target protein. This latter issue, known as a ‘linkage error’, particularly impacts accuracy and precision when using super-resolution techniques. The flexibility of the antibody scaffold also means that the distance between the fluorophore and the target is often random, and therefore challenging to correct. These limitations have driven the development of smaller probes, such as Affimers and nanobodies.
Michelle Peckham and Darren Tomlinson are group leaders at the University of Leeds. Together, their research teams have been developing a range of Affimers, small alternatives to antibodies that are less than 4 nm long. In a new Review for Journal of Cell Science (JCS), the Peckham and Tomlinson labs explain the advantages that Affimers offer over traditional labelling techniques, often using examples from their own work. Glenn Carrington is a postdoctoral researcher in the Peckham lab and a co-author on the Review. “With the advent of super-resolution imaging technologies, the need for specific high-affinity small binding proteins has increased,” Glenn explained. “Affimers and nanobodies fill this role seamlessly.” Here, we highlight some of their discussion. For further information, head to issue 14 of JCS to check out the full review.
What is an Affimer?
Affimers are small, engineered binding proteins whose scaffold is based on the consensus sequence of either human stefin A (type I Affimers) or plant phytocystatin (type II Affimers), two proteins that act as cysteine protease inhibitors. They also have a binding interface comprising two short regions of variable sequence, and it is this part of the Affimer that interacts with your protein of interest.
Besides their small size and stability, the authors point out that Affimers offer the added advantage of being relatively quick to produce. Screening for Affimers that bind your protein of interest is carried out in vitro, using a phage display approach, and can typically be completed within two weeks. This is clearly much faster than traditional antibody or nanobody production methods, which usually require you to inoculate an animal with your target protein.
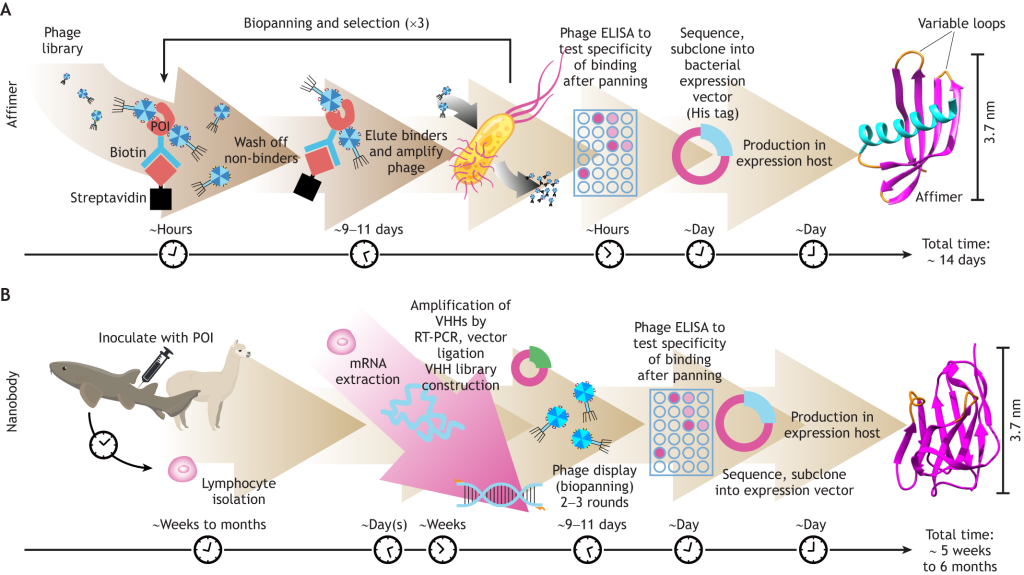
Fluorescence imaging applications
To prepare Affimers for imaging applications, the authors explain that their favoured approach is maleimide bioconjugation chemistry. In other words, they make use of a single cysteine residue, which they introduce at either the N- or C-terminus of the Affimer. This enables them to couple their fluorescent dye of choice to the Affimer.
The authors go on to present a series of case studies demonstrating how they and other groups have used Affimers in their imaging experiments. “In my research, Affimers are proving robust and flexible reagents for super-resolution microscopy, to image protein modifications in dividing cells and to gain insights into the structure of protein complexes in MAP kinase signalling,” co-author Paul Cordell told us. “I was interested to compare Affimers with nanobodies, to learn more about their respective strengths and weaknesses, especially in imaging applications.” Like nanobodies, Affimers can be used in both fixed and live cells. We summarise a few interesting example below:
- Fixed cell imaging
The authors have previously generated Affimers that can label filamentous actin (F-actin) in cells fixed with methanol. This is a great example of how Affimers can offer viable alternatives to some of the limitations of traditional methods, since phalloidin, a widely used actin stain, does not work under methanol fixation conditions. - Live cell imaging
The authors have also used Affimers to image actin in live cells, with an eGFP molecule connected to the Affimer using a flexible linker. They also discuss the ease with which Affimers can be adapted for different applications; for example, Affimers that are coupled to GFP via rigid linkers are suitable for fluorescence polarisation microscopy, and have been used to investigate the orientation of F-actin filaments in live cells. - Combining Affimers with super-resolution imaging
The authors explain that the sheer size of traditional antibody probes is a limiting factor when optimising the accuracy of super-resolution imaging. Using an example from their own work, they recount how they were able to use an Affimer to visualise Z-discs with 3D dSTORM. As well as being able to penetrate the dense structure of the Z-discs, the Affimer allowed them to measure the spacing of α-actinin-2. Their measurement of 18.5 nm was close to that reported via electron microscopy.
In summary
Antibodies are an effective tool for labelling proteins of interest, but their large size is making them increasingly incompatible with super-resolution imaging techniques. Small probes like Affimers and nanobodies offer an alternative, and this Review is a great introduction if you are considering incorporating these tools in your own imaging experiments. Reflecting on this changing landscape of labelling techniques, the authors state that they “anticipate that the use of these small probes is likely to increase exponentially in the future, and that they will become the standard tools for super-resolution imaging”.
Read the full Review on JCS:
Affimers and nanobodies as molecular probes and their applications in imaging


 (No Ratings Yet)
(No Ratings Yet)