Back to the density: using Surface Morphometrics to measure organelle membrane thickness in cryo-electron tomography
Posted by Michaela Medina, on 18 March 2026
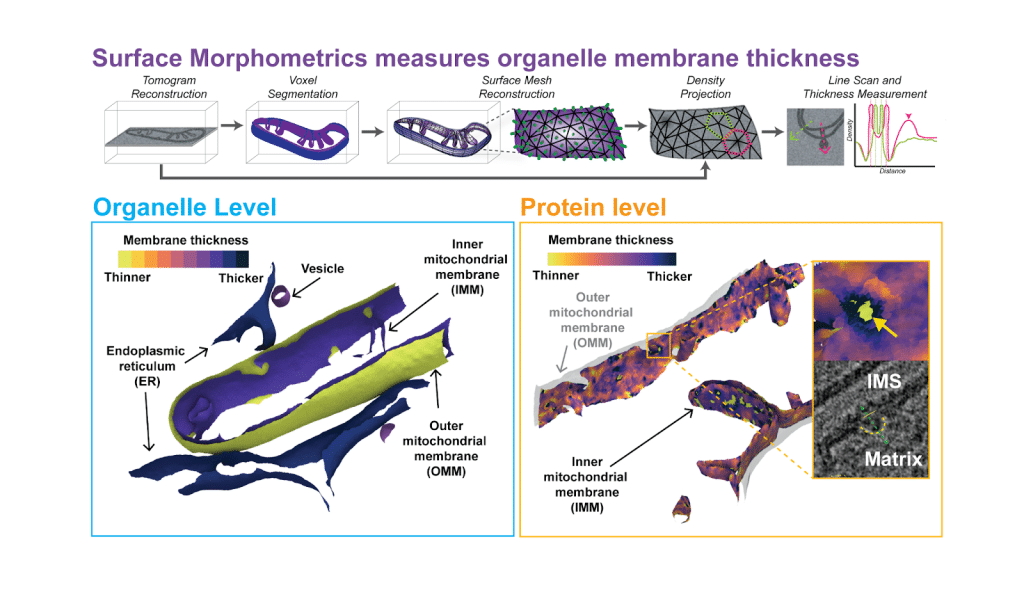
Surface Morphometrics is an open-access software that extracts quantitative information of membrane ultrastructure from high resolution cryo-electron tomography (cryo-ET) data. This tooling allows for the study of organelles in their native environment across large populations in an automated fashion. The Surface Morphometrics pipeline reconstructs three-dimensional triangular surface meshes to model and quantify the cellular membrane ultrastructure from cryo-electron tomography (cryo-ET) data. The initial version of Surface Morphometrics enabled detailed measurements of membrane geometry, including inter-/intra membrane distances, curvedness, and relative orientation between different cellular membranes. In our most recent publication, we extend this pipeline to directly measure membrane thickness in cells- a measurement that was previously possible through in vitro systems. This is achieved by surface mesh reconstructions to guide voxel-density line scans across organellar membranes in cryo-ET data.
How has it been used so far?
Within our publication we show that distinct organelles have distinct membrane thicknesses, a feature conserved in mammalian and yeast cells. We further reveal that membranes with distinct regions such as the inner mitochondrial membrane have different thicknesses. Zooming in further we looked at the local membrane environment and found that extreme variations in membrane thickness correlate with large macromolecular protein complexes.
Excitingly we have applied the expanded surface morphometrics pipeline to investigate organelle membranes in different biological systems such as flavivirus replication and maturation in host endoplasmic reticulum membranes (1) and amyloid induced lysosomal membrane damage (2). Additionally this tooling was used to interrogate local membrane environments around large macromolecular complexes within mitochondrial membrane such as ATP synthase and prohibitin complexes (3).
The power of surface morphometrics lies in its ability to unify diverse aspects of membrane structure into a single quantitative framework, enabling comparisons across properties such as curvedness, distance, orientation – and now thickness! Specificity of membrane analysis can be achieved through membrane- and protein- feature filtering strategies, allowing for targeted exploration of ultrastructural characteristics. We are excited to see how the community will leverage this tooling to probe new facets of cell biology and reveal novel mechanistic insights.
- Dahmane, S., Schexnaydre, E., Zhang, J. et al. Cryo-electron tomography reveals coupled flavivirus replication, budding and maturation. Nat Commun 17, 828 (2026). https://doi.org/10.1038/s41467-026-68483-4
- Li, D., Zhang, W. et al. Cathepsin-dependent amyloid formation drives mechanical rupture of lysosomal membranes. bioRxiv. (2026). https://doi.org/10.64898/2026.01.17.700056
- Medina, M. et al. Prohibitin complexes associate with unique membrane microdomains in cells. bioRxiv. (2026). https://doi.org/10.1101/2025.11.14.688579
What do you need to be able to use it?
Users will need three-dimensional tomographic data in which the lipid bilayer of organelle membranes is resolved, along with voxel segmentation of membrane. Surface Morphometrics can be installed by following the instruction provided in our Github repository (https://github.com/GrotjahnLab/surface_morphometrics). To visualize the quantification result on surface meshes generated by Surface Morphometrics, user will need to install Paraview or the recently developed Surface Morphometrics GUI from Barad lab in OHSU (https://github.com/baradlab/surface-morphometrics-gui).
What are the future plans for further development?
Currently, studying the local membrane environment of proteins still largely relies on user-defined selections of specific proteins. A key future direction will be to make this process more robust and unbiased by integrating unsupervised screening of cellular membrane surfaces with clustering of geometric features produced by the Surface Morphometrics pipeline. Such approaches would enable systematic identification of recurring membrane geometries, opening the door to visualizing previously invisible aspects of the cellular proteome and revealing functionally relevant subdomains of organelles directly in their native cellular context.
More information
You can check out our publication here for more methodological details: Michaela Medina, Ya-Ting Chang, Hamidreza Rahmani, Mark Frank, Zidan Khan, Daniel Fuentes, Frederick A. Heberle, M. Neal Waxham, Benjamin A. Barad, Danielle A. Grotjahn; Surface Morphometrics reveals local membrane thickness variation in organellar subcompartments. J Cell Biol 2 March 2026; 225 (3): e202505059. doi: https://doi.org/10.1083/jcb.202505059
Further news and updates for the software can be found in our Github: https://github.com/GrotjahnLab/surface_morphometrics
The tomograms used in this study are open access and can be found in Electron Microscopy Public Image Archive under accession code EMPIAR-12534


 (1 votes, average: 1.00 out of 1)
(1 votes, average: 1.00 out of 1)