Featured image with Alexis Lomakin
Posted by FocalPlane, on 22 March 2023
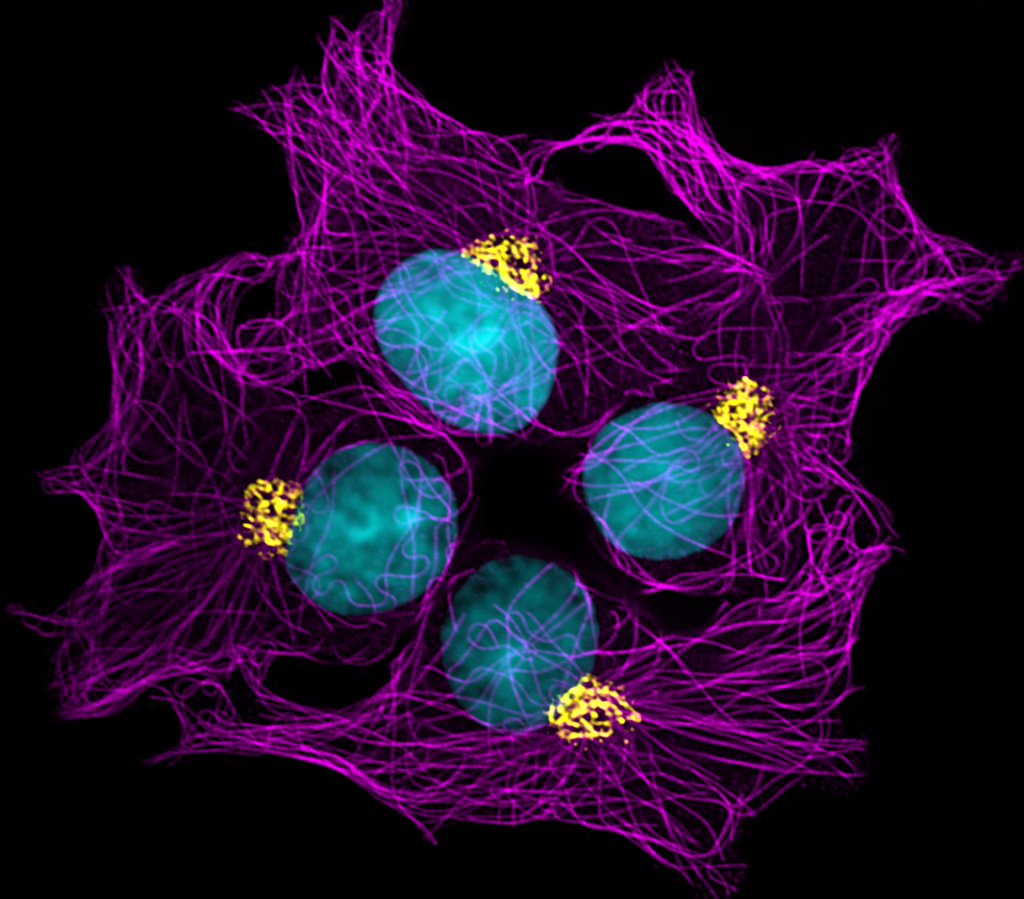
Our featured image, ‘Cellular highways’ shows a small group of rat liver epithelial cells IAR-2 and their internal organelles: the nucleus (cyan), cytoskeletal microtubules (magenta) and the Golgi apparatus (yellow). The microtubules form ‘cellular highways’ that mediate protein and lipid transport via cytoplasmic vesicles such as those that participate in the biogenesis and dynamics of the Golgi.
Except for the nucleus whose chromatin was directly labelled with the blue-fluorescent DNA stain DAPI, these structures were revealed using indirect immunofluorescence labelling in which ice-cold methanol fixed-permeabilised cells were exposed to primary antibodies recognising protein markers of microtubules (alpha-tubulin) and the Golgi apparatus (giantin). Species-specific secondary antibodies conjugated with spectrally non-overlapping fluorophores were subsequently utilised to highlight the primary antibodies and thus the proteins markers of interest.
The sample was imaged using Yokogawa spinning disk confocal system. The confocal z-stack was then pseudocoloured, processed in MetaMorph and represented as a single, maximum-intensity projection image.
We caught up with Alexis Lomakin to find out more about the research behind the image.
Research career so far: The image was taken while I was a postdoc at Harvard Medical School (Boston, MA, USA) in the laboratory of Gaudenz Danuser. I now run my own laboratory at the Medical University of Vienna (Vienna, Austria).
Current research: As follows from the image, I have a long-standing interest in the fundamental question of how epithelial tissue cells organise themselves in space-and-time and how they translate this organisation into a specific function in physiological and pathophysiological situations. Currently, my group seeks to quantitatively elucidate biochemical signalling and physicochemical mechanisms allowing cells to scale the rate of metabolic reactions to intracellular organelle abundance and size. We are particularly interested in how these mechanisms become dysregulated in human epithelial tumour cells that develop tolerance towards chemotherapeutic stresses triggering cellular aging.
Favourite microscope/imaging technique: I am a big fan of compact benchtop microscopes for automated high-content fluorescence imaging that you can have right next to your bench where the actual experiment is being done. These systems do not necessarily have to be confocal or ‘super-duper’ resolution, but they should have high-quality optics and cameras along with a broad range of filters to allow multispectral imaging. Coupled with a powerful computer, these types of microscopes are great at capturing massive amounts of microscopy data in a highly controlled and consistent fashion, something that is critical when it comes to quantitative multiparametric image analyses.
In low-throughput imaging experiments, I prefer capturing images of single cells using a high numerical aperture high magnification lens on a laser confocal microscope.
What are you excited about in microscopy: I am excited about the emerging tools based on machine and deep learning for the processing and analysis of microscopy data. The efforts to democratise them, to make them publicly available and user-friendly, are invaluable. I am also happy to see commercial microscopy analysis products being designed in a way amenable to the integration with publicly available tools; this is something that my lab is currently trying to pursue in a collaboration with Thermo Fisher Scientific Inc. We are keen on these analytical tools because we strive to measure cellular and subcellular traits in each and every cell to quantitatively understand the observable phenotypic heterogeneity at a single-cell level. Indeed, ´seeing is believing´, is something that single-cell molecular -omics techniques could not offer.


 (6 votes, average: 1.00 out of 1)
(6 votes, average: 1.00 out of 1)