Microscopy preprints: bioimage analysis
Posted by FocalPlane, on 20 March 2026
Here is a curated selection of preprints published (or updated) recently. In this post we focus specifically on bioimage analysis and data management.
Packaging Jupyter notebooks as installable desktop apps using LabConstrictor
Iván Hidalgo-Cenalmor, Marcela Xiomara Rivera Pineda, Bruno M. Saraiva, Ricardo Henriques, Guillaume Jacquemet
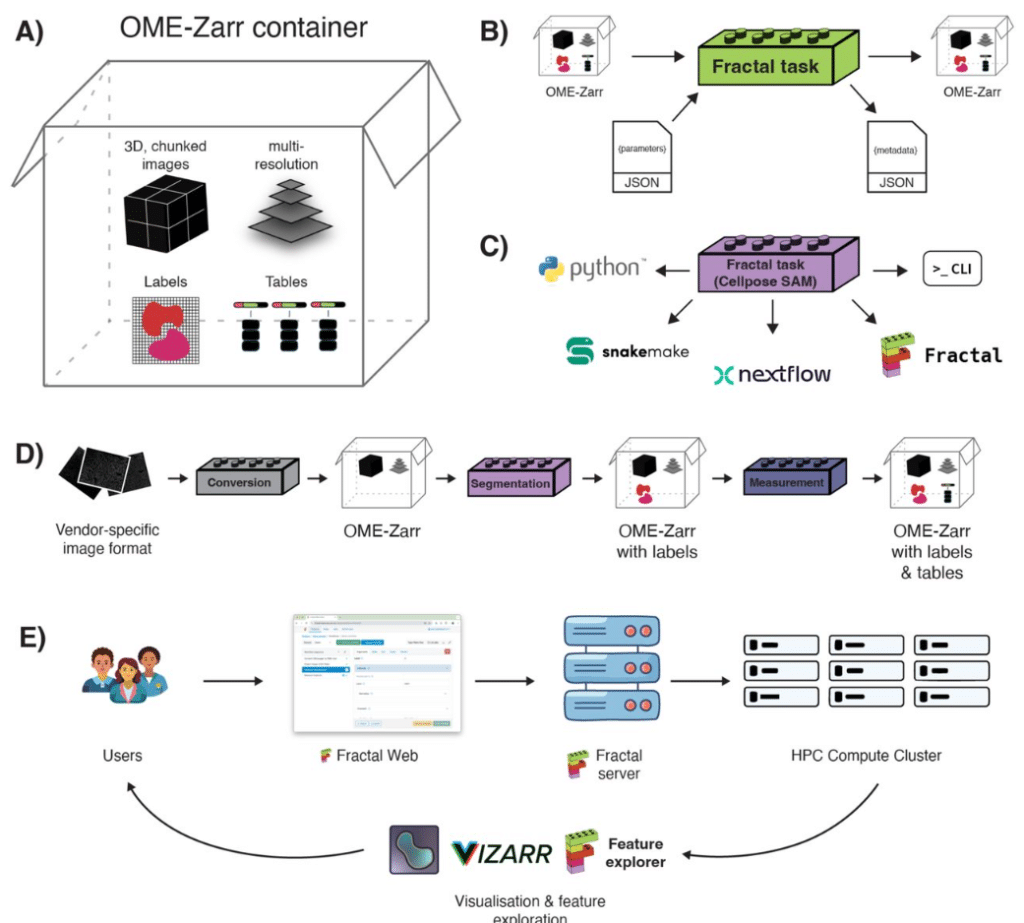
Fractal: Towards FAIR bioimage analysis at scale with OME-Zarr-native workflows
Joel Lüthi, Lorenzo Cerrone, Tommaso Comparin, Max Hess, Ruth Hornbachner, Adrian Tschan, Gustavo Quintas Glasner de Medeiros, Nicole A. Repina, Loredana K. Cantoni, Fabio D. Steffen, Jean-Pierre Bourquin, Prisca Liberali, Lucas Pelkmans, Virginie Uhlmann
A general methodology for liver sinusoid fenestration analysis based on 3D electron microscopy data
Cécile Pohar, Yousr Rekik, Minh Son Phan, Benoit Gallet, Agnès Desroches-Castan, Mireille Chevallet, Jean-Yves Tinevez, Emmanuelle Tillet, Nicola Vigano, Pierre-Henri Jouneau, Aurélien Deniaud
MuViT: Multi-Resolution Vision Transformers for Learning Across Scales in Microscopy
Albert Dominguez Mantes, Gioele La Manno, Martin Weigert
High-Fidelity Long-term Whole-embryo Lineage and Fate Reconstruction by Iterative Tracking with Error Correction
Mengfan Wang, Qinghua Zhang, Congchao Wang, Yunfeng Chi, Wei Zheng, Zeyu Mu, Xiangyu Cao, Weizhan Zhang, Boao Yang, Alexander F. Schier, Joaquin Navajas Acedo, Yinan Wan, Guoqiang Yu
Phasor analysis of RGB camera data enables fluorescence microscopy unmixing and brightfield segmentation in a commercial microscope
Bruno Schuty, María José García, Satya Khuon, Leonel Malacrida
Interactive segmentation of membrane and membrane mimic densities in cryo-EM maps
Alok Bharadwaj, Lotte Veerbeek, Arjen J. Jakobi
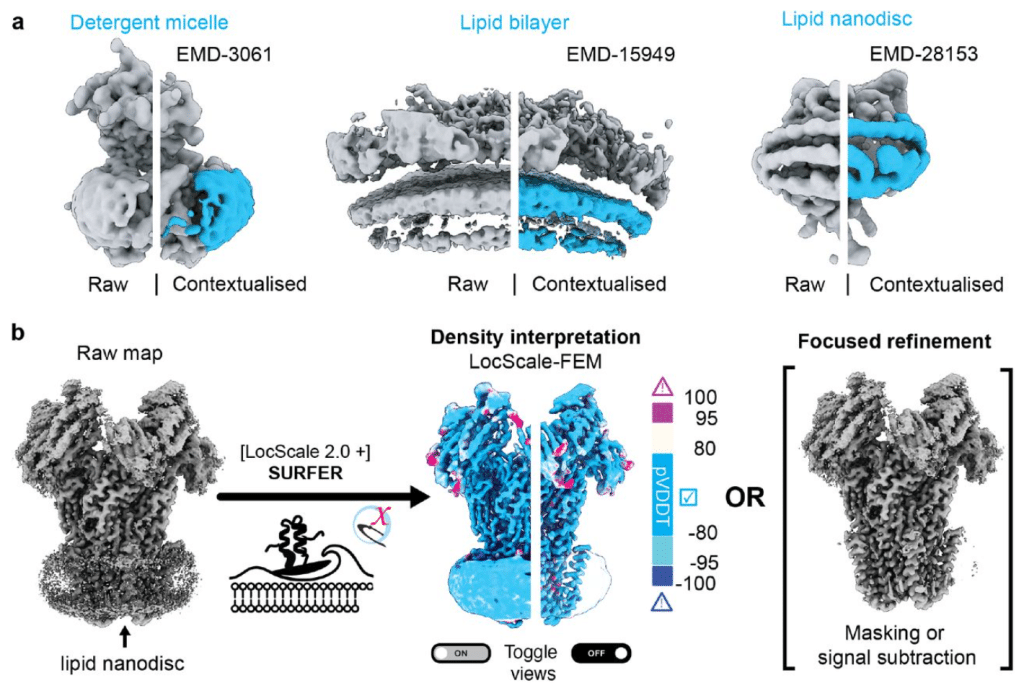
Image Analysis Tools for Electron Microscopy
David H. Shtengel, Gleb Shtengel, C. Shan Xu, Harald F. Hess
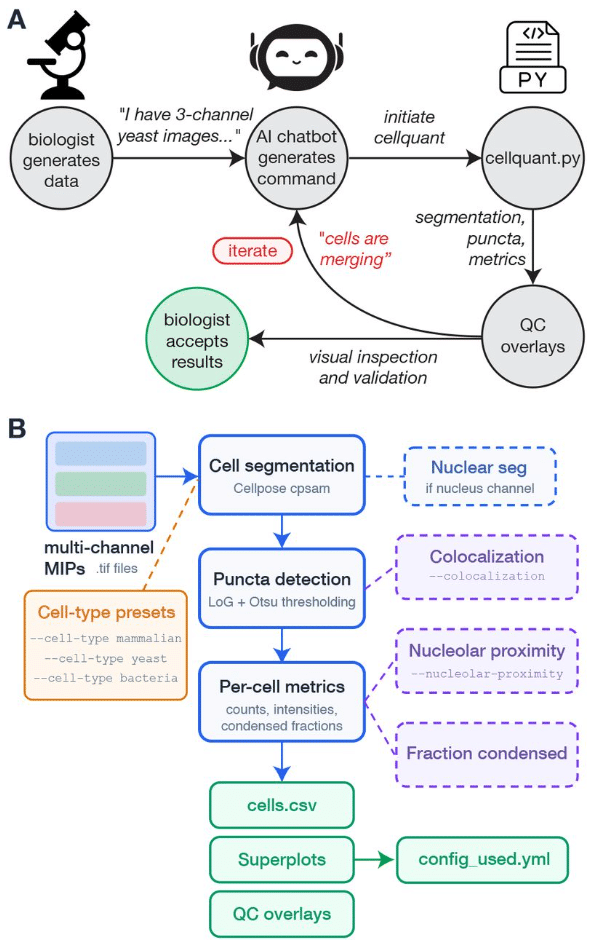
Cellquant: a vibecoder’s guide to image analysis
Abani Neferkara, Asif Ali, David Pincus
A foundation AI model enhances electron microscopy image analysis
Mengze Du, Yuwei Wang, Li Xie, Gaoliang Deng, Jiansheng Guo, Bo Han, Zhong-Hua Chen, Chen Rui, Jiankang Han, Yuan Chen, Yanru Zhao, Runzhou Cao, Fei Wang, Kun Li, Yu Wang, Yong He, Xuping Feng
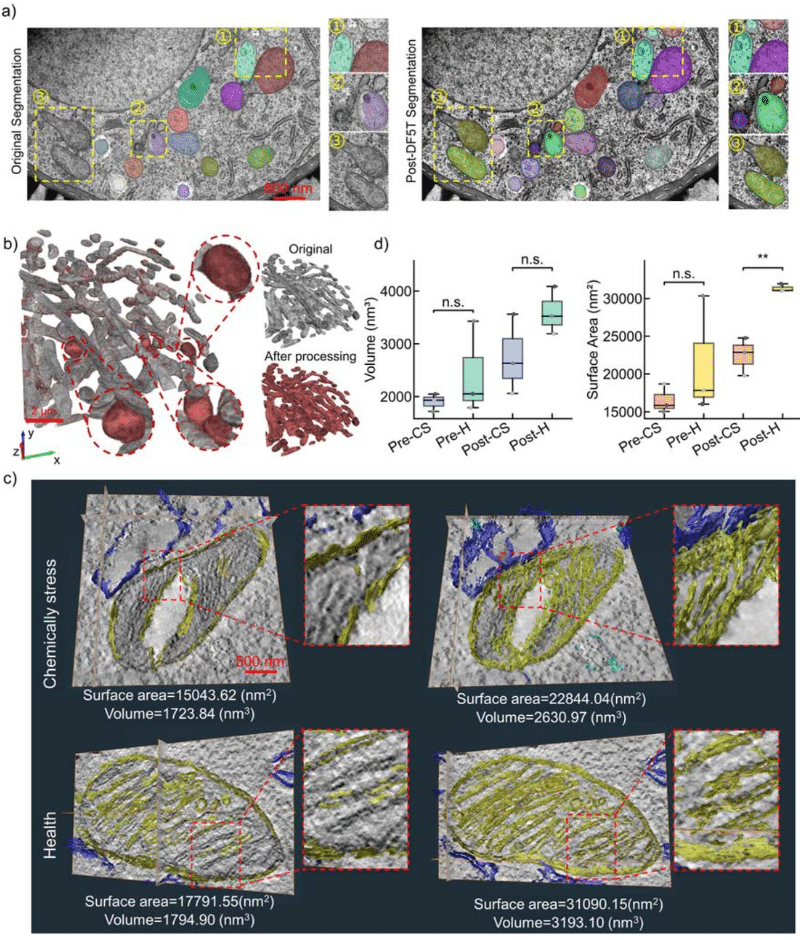
Benchmarking Machine Learning and Automated Image Analysis for Organelle Quantification
Chloé Daul, Pierre Tournier, Shukry J. Habib
SynAPSeg: A novel dataset and image analysis framework for deep learning-based synapse detection and quantification
Pascal Schamber, Sahana Darbhamulla, Molly Boyer, Madison Pelletier, Helene Hartman, Olivia Friedman, Shiyu Zhang, Allison Blais, Seyun Oh, Haining Zhong, Alexei M Bygrave
Evaluating image upsampling strategies for downstream microscopy image classification
Sakib Mohammad, Aalvee Asad Kausani, Md Noumil Tousif
Cryo-EM image processing of amyloid filaments in RELION-5.1
Sofia Lövestam, Jenny Shi, David Li, Kiarash Jamali, Sjors H.W. Scheres
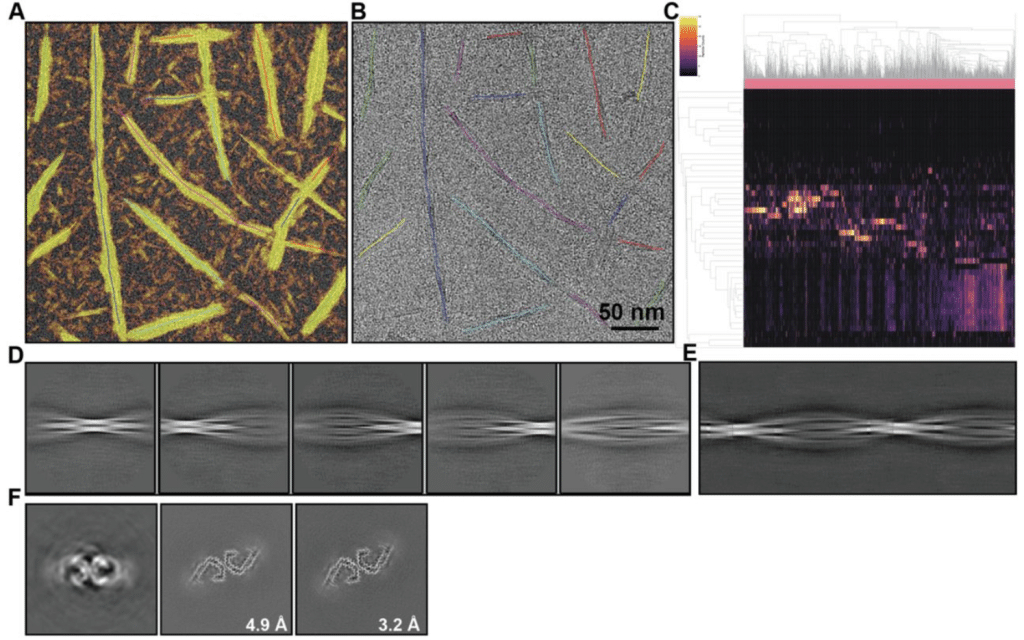
cryoJAX: A Cryo-electron Microscopy Image Simulation Library In JAX
Michael J. O’Brien, David Silva-Sánchez, Geoffrey Woollard, Kwanghwi Je, Sonya M. Hanson, Daniel J. Needleman, Pilar Cossio, Erik Henning Thiede, Miro A. Astore
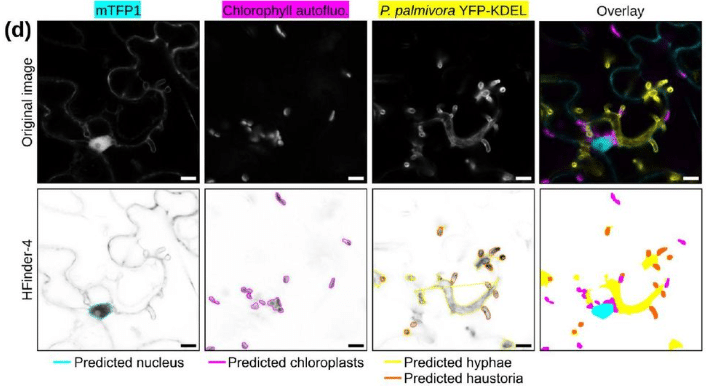
Deep learning enables quantitative subcellular analysis of plant-microbe interfaces
Stavros Korovesis, Shumei Wang, Lin Xu, Inès Giraudon, Daniela Rosales Hernandez, Enora Panek, Laure Boeglin, Maria-Myrto Kostareli, Max HJ Pluis, Baoyu Wang, Yuan Wang, Djenatte Abdennour, Harald Keller, Paul RJ Birch, Sebastian Schornack, Edouard Evangelisti
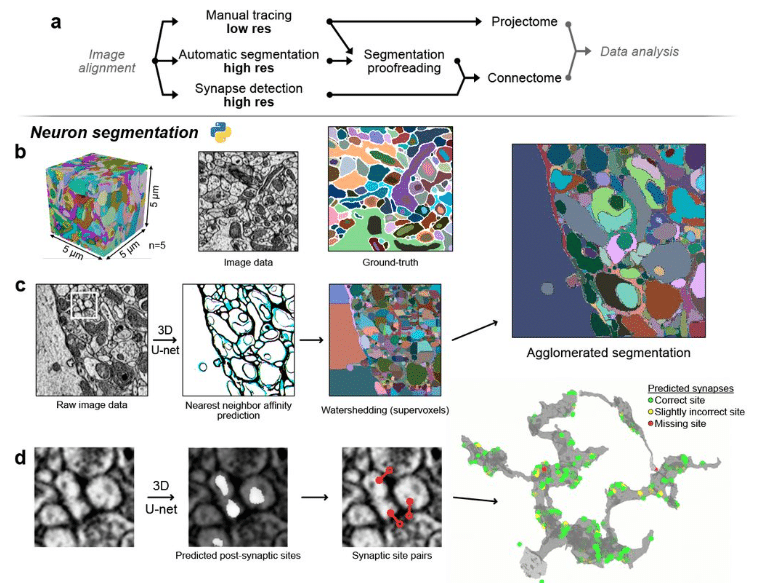
A multi-resolution imaging and analysis pipeline for comparative circuit reconstruction in insects
Valentin Gillet, Marcel E. Sayre, Griffin S. Badalamente, Nicole L. Schieber, Kevin Tedore, Jan Funke, Stanley Heinze
A Modular Framework for Automated Segmentation and Analysis of AFM Imaging of Chromatin Organization
Emily Winther Sørensen, Sushil Pangeni, Raquel Urteaga-Merino, Peter J. Murray, Sergei Rudnizky, Ting-Wei Liao, Fahad Rashid, Jihee Hwang, Maryam Yamadi, Xinyu A. Feng, Jonas Zähringer, Stephanie Gu, Iain F. Davidson, Laura Caccianini, Manuel Osorio-Valeriano, Lucas Farnung, Seychelle M. Vos, Jan-Michael Peters, James Berger, Carl Wu, Nikos S. Hatzakis, Julius B. Kirkegaard, Taekjip Ha
An Integrated and Configurable End-to-End Pipeline for Longitudinal Cell Painting Analysis
Guang Zhao, Yu Xi, Mikhail Titov, Rebecca Weinberg, Sara Forrester, Tom Brettin, Shinjae Yoo, Xiaoning Qian, Byung-Jun Yoon
ExoFILT: Transfer learning for robust and accelerated analysis of exocytosis single-particle tracking data
Eric Kramer, Laura I. Betancur, Sasha Meek, Sébastien Tosi, Carlo Manzo, Baldo Oliva, Oriol Gallego
The One Click Wonder: a retrained automated segmentation pipeline that enables quantitative and modular analysis of C. elegans embryos
Palmer Carlyn Bassett, Tobias Evan Verheijen, Angelo Luiz Angonezi, Aude Andriollo, Sebastien Herbert, Gregory Roth, Jeffrey A. Chao, Susan E. Mango
NucVerse3D: Generalizable 3D nuclear instance segmentation across heterogeneous microscopy modalities
Jorge Vergara, Cristian Perez-Gallardo, Ricardo Velasco, Dilan Martinez, Diego Badilla, Esteban G. Contreras, Pamela Guevara, Fabián Segovia-Miranda, Hernán Morales-Navarrete
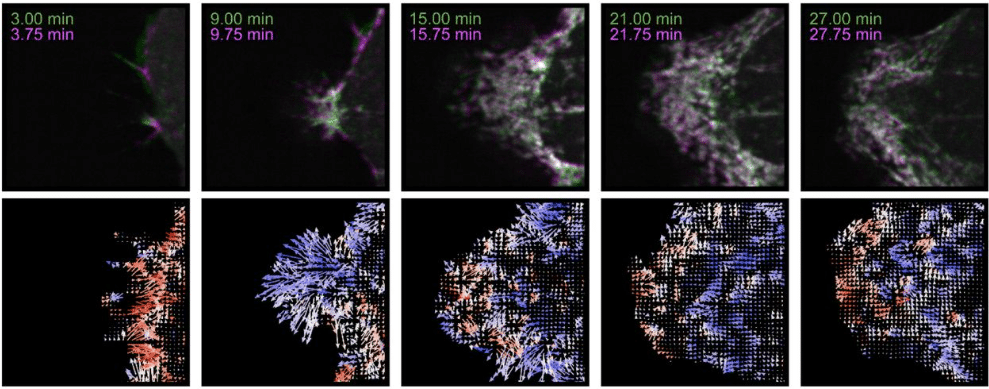
Measuring Amorphous Motion: Application of Optical Flow to Three-Dimensional Fluorescence Microscopy Images
Rachel M. Lee, Leanna R. Eisenman, Chad M. Hobson, Jesse. S. Aaron, Teng-Leong Chew
Inverse Protocol Prediction from Spheroid Microscopy Imaging via Morphology-Aware Structured Learning
Prateek Mittal, Ayush Srivastava, Joohi Chauhan
BEEP Learning: Multi-View Image Decomposition for Massively Multiplexed Biological Fluorescence Microscopy
Ruogu Wang, Thet Teresa Hnin, Yunlong Feng, Alex M. Valm
A semantic segmentation model to predict subcellular glycogen localization using transmission electron microscopy images
Anders A. Hansen, Jacob M. Egebjerg, Kristian Solem, Kristoffer J. Kolnes, Daniel Wüstner, Jørgen F. P. Wojtaszewski, Jørgen Jensen, Joachim Nielsen


 (1 votes, average: 1.00 out of 1)
(1 votes, average: 1.00 out of 1)