Microscopy preprints: bioimage analysis
Posted by FocalPlane, on 1 May 2026
Here is a curated selection of preprints published (or updated) recently. In this post we focus specifically on bioimage analysis and data management.
TomoSwin3D: a Swin3D Transformer for the Identification and Classification of Macromolecules in 3D Cryo-ET Tomograms
Ashwin Dhakal, Rajan Gyawali, Jianlin Cheng
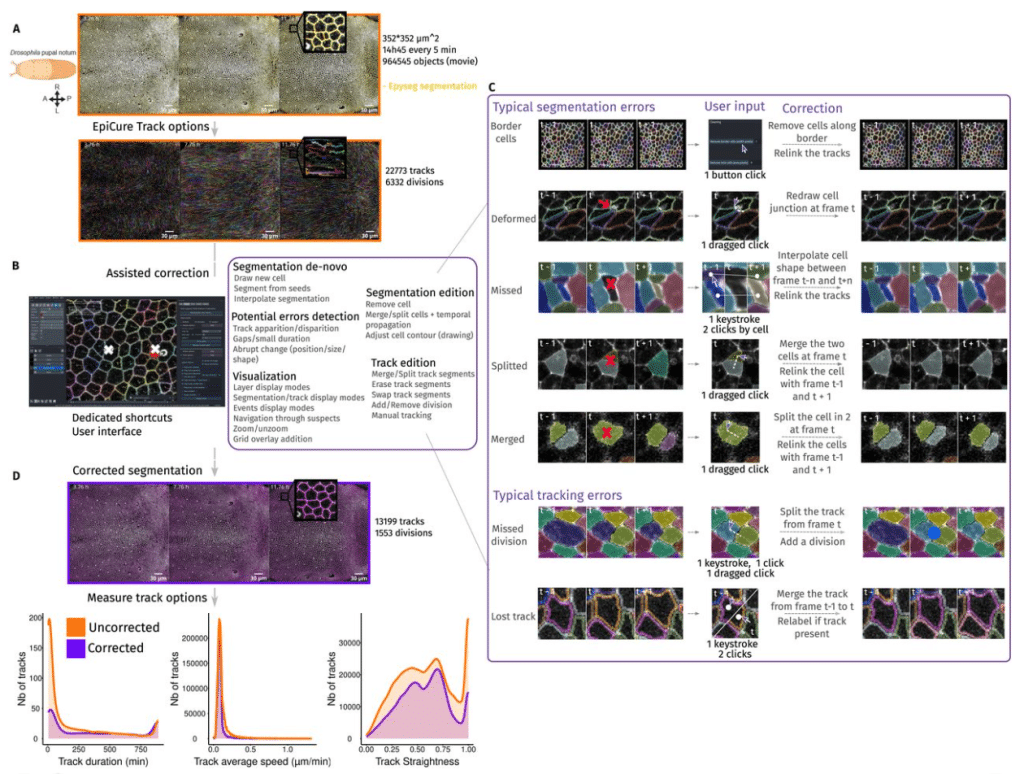
EpiCure (Epithelial Curation): a versatile and handy tool for curation of epithelial segmentation
Gaëlle Letort, Léo Valon, Arthur Michaut, Tom Cumming, Laura Xénard, Minh-Son Phan, Nicolas Dray, Curtis T Rueden, François Schweisguth, Jérôme Gros, Laure Bally-Cuif, Jean-Yves Tinevez, Romain Levayer
Shape2Fate: a morphology-aware deep learning framework for tracking endocytic and exocytic carriers at nanoscale
Adam Harmanec, Alexander D. Dagg, Jan Kamenicky, Tomas Kerepecky, Yelyzaveta Makieieva, Conceição Pereira, Nicholas Bright, Dilip Menon, David C. Gershlick, Nadezda Vaskovicova, Tiffany Lai, Daniel Fazakerley, Lothar Schermelleh, Filip Sroubek, Zuzana Kadlecova
Image Analysis Tools for Scanning Electron Microscopy
David H. Shtengel, Gleb Shtengel, C. Shan Xu, Harald F. Hess
Quantitative Mapping of Organelle Positioning in Cultured Cells Using Semi-Automated Image Analysis Pipeline
Katerina Jerabkova-Roda, Vincent Hyenne, Jacky G. Goetz
A Decade of Deep Learning-based Biomedical Image Segmentation
Suhao Yu, Haojin Wang, Ningsen Wang, Sicheng Chen, Juncheng Wu, Zhenlong Yuan, Tianhao Qi, Zongwei Zhou, Fei Xia, Jun Ma, Yuyin Zhou
CRISP enables comparisons of image-based spatial transcriptomic segmentation quality across ten organs
James R. Rose, Elex S. Rose, José A. F. Assumpção, Heather Pathak, Hannah E. Peck, Loren E. Sasser, Charmy J. Patel, Daryll Vanover, Philip J. Santangelo
cryoAgent: An agentic workflow for robust and adaptive end-to-end cryo-EM image processing
Daoyi Li, Shaodi Yang, Qian Xiao, Tongxin Niu, Yan Zhang, Yun Zhu, Fei Sun
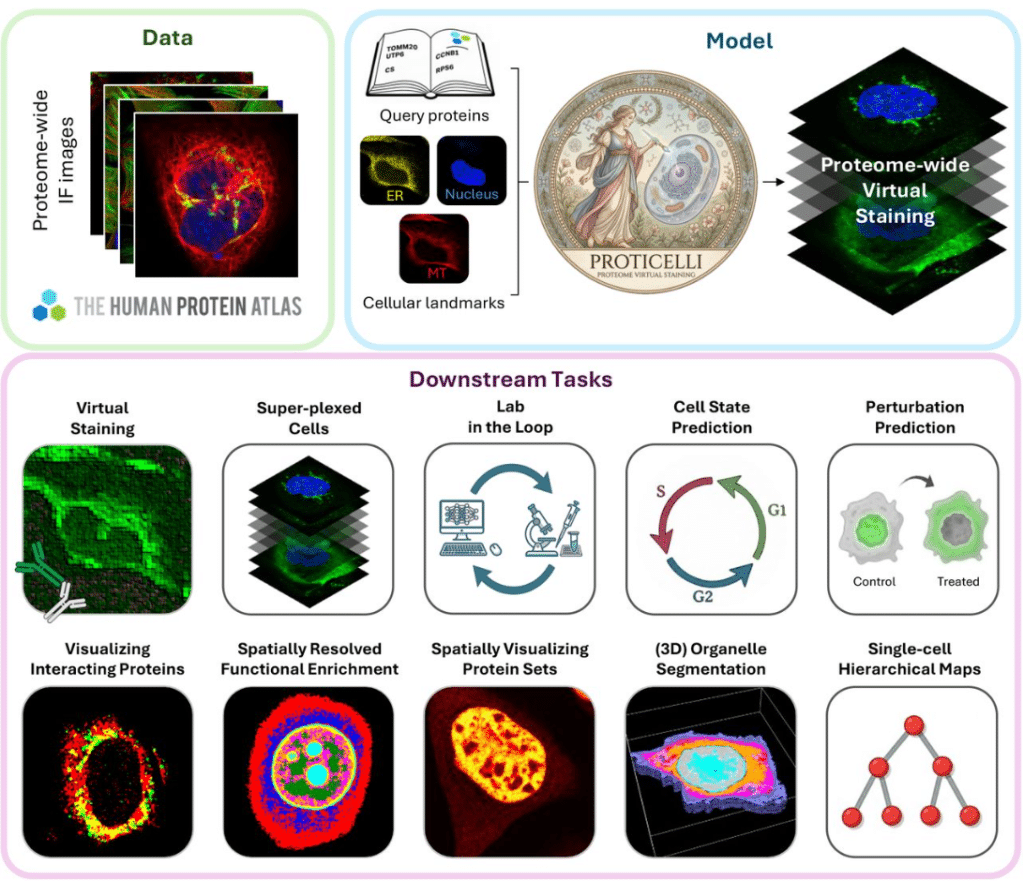
Generative machine learning unlocks the first proteome-wide image of human cells
Huangqingbo Sun, Konstantin Kahnert, Jan N. Hansen, William Leineweber, Mingyang Li, Wanyue Feng, Frederic Ballllosera, Ulrika Axelsson, Wei Ouyang, Emma Lundberg
Benchmarking three simple DNA staining-based image metrics for live-cell tracking of chromatin organization
Minwoo Kang, Aidan Tomas Cabral, Manasi Sawant, Hawa Racine Thiam
DCAFA: Differential Community Abundance and Feature Analysis for Histological Images
George Wright, Piotr Keller, Joanne Muter, Jan Brosens, Sabine Tejpar, Fayyaz Minhas
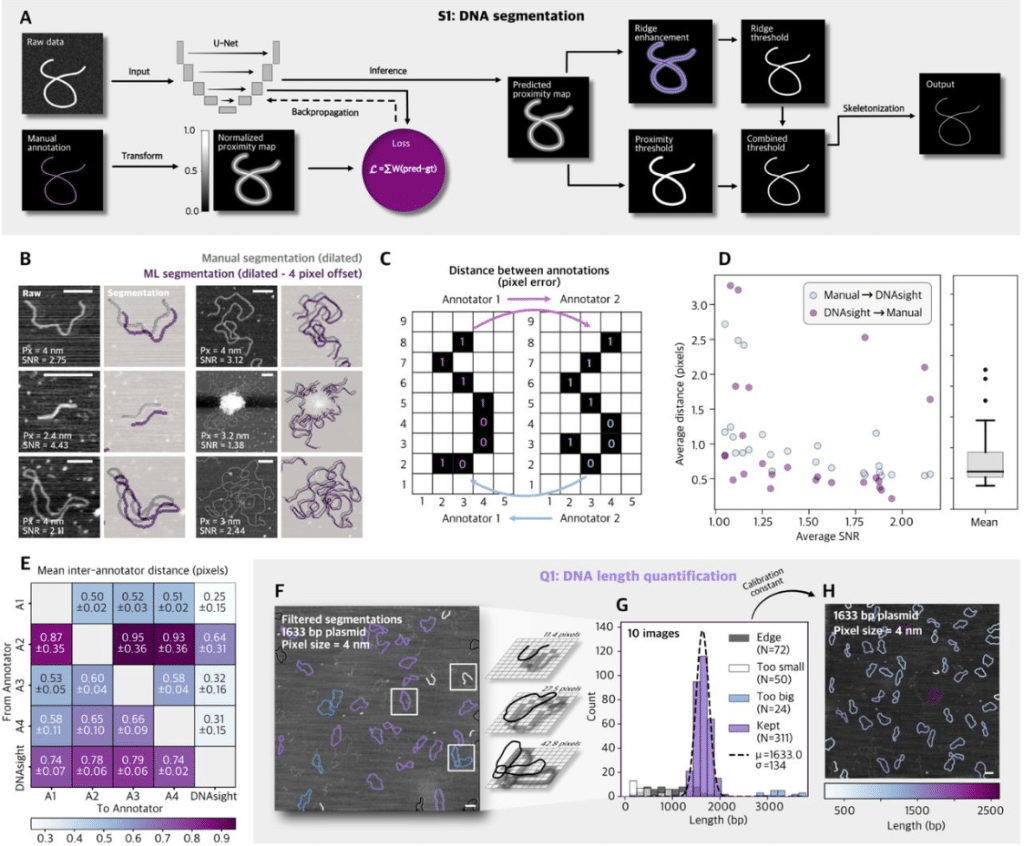
A Modular Framework for Automated Segmentation and Analysis of AFM Imaging of Chromatin Organization
Emily Winther Sørensen, Sushil Pangeni, Raquel Urteaga-Merino, Peter J. Murray, Sergei Rudnizky, Ting-Wei Liao, Fahad Rashid, Jihee Hwang, Maryam Yamadi, Xinyu A. Feng, Jonas Zähringer, Stephanie Gu, Iain F. Davidson, Laura Caccianini, Manuel Osorio-Valeriano, Lucas Farnung, Seychelle M. Vos, Jan-Michael Peters, James Berger, Carl Wu, Nikos S. Hatzakis, Julius B. Kirkegaard, Taekjip Ha
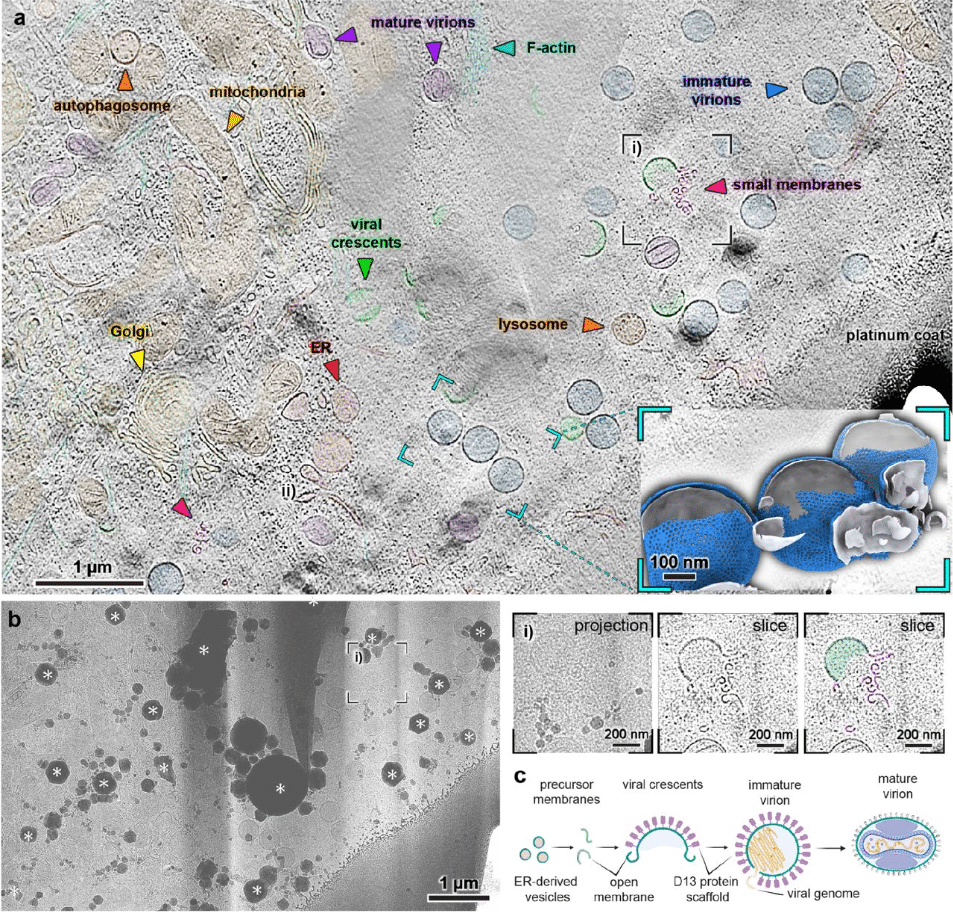
View Tomo: Context-aware targeting and analysis in electron cryo-tomography
Robert Gebauer, Emily A. Machala, Yuliia Mironova, Märit-Runa Jönsson, Jędrzej Mazur, Clara A. Feldmann, Marcel A. Zimmeck, Emma Silvester, Enrico Caragliano, Julian Falckenhayn, Enoch Lok Him Yuen, Tarhan Ibrahim, Jan Hellert, Tolga O. Bozkurt, Rainer Kaufmann, Emmanuelle R. J. Quemin, Kay Grünewald, Vojtěch Pražák
Anchored Brownian motion and Bayesian methods for the analysis of single particle tracking data
Alan Boles, Robert J. Pyron, Sarah Shelby, Ioannis Sgouralis
Cilia SubQ, a modular suite of pipelines for automated analysis of primary cilia and ciliary subdomains
Elizabeth Menzel, Karama Hamdi, Gracie Hoffman, Abdelhalim Loukil
How Mitochondria Distribute and Align in HeLa Cells: A Computational Analysis On Electron Microscopy Images
Daniel Brito-Pacheco, Panos Giannopoulos, Constantino Carlos Reyes-Aldasoro
NetTracer3D Enables User-Friendly Analysis of Diverse Microscopic and Medical 3D Datasets
Liam Mclaughlin, Mia Curic, Siddhartha Sharma, Jorge Villazon, Rebecca J. Salamon, Manako Yamaguch, Maria Luisa S. Sequeira-Lopez, Philippa R. Kennedy, Riley C. Lyons, Lingyan Shi, R. Ariel Gomez, Sanjay Jain
Label-free 4D holotomography with depth-adaptive segmentation for quantitative analysis of lipid droplet dynamics in hepatic organoids
Jimin Cho, Hoyeon Lee, ChulMin Oh, Juyeon Park, Sujin Park, Bon-Kyoung Koo, YongKeun Park
Semi supervised GAN for smart microscopy, fast and data efficient cell cycle classification
Rajeev Manick, Youssef El Habouz, Maëlle Guillout, Celia Martin, Julia Bonnet, Louis Ruel, Sylvain Pastezeur, Olivier Chanteux, Otmane Bouchareb, Marc Tramier, Jacques Pécréaux
A novel attention mechanism for noise-adaptive and robust segmentation of microtubules in microscopy images
Achraf Ait Laydi, Louis Cueff, Mewen Crespo, Yousef El Mourabit, Hélène Bouvrais
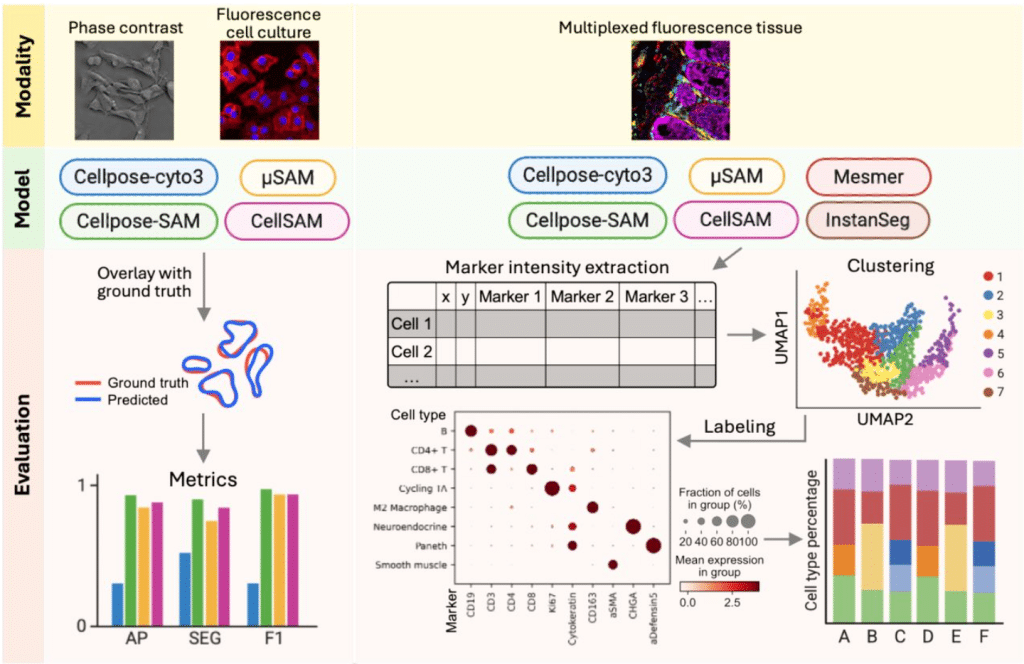
Foundation cell segmentation models performance on live microscopy and spatial-omics data
Yang Miao, Nick Surguladze, Josh Lerner, Koravit Poysungnoen, Ky Ariano, Yuexi Li, Yining Zhu, Kyra Van Batavia, Jodie Jepson, Josie Van De Klashorst, Bobby Y.X. Ni, Alexander Armstrong, Reeha Rahman, Roarke Horstmeyer, John W. Hickey
HYPER-Net: Physics-Conditioned Self-Supervised Reconstruction for Fourier Light-Field Microscopy
Zhi Ling, Xuanwen Hua, Wenhao Liu, Hao Wu, Lucinda M Peng, Jessica Hou, Parvin Forghani, Christopher Pierce, Ge-Ah R Kim, Shuichi Takayama, Shuyi Nie, Chunhui Xu, Hang Lu, Shu Jia
Deep Learning Enables Automated Segmentation and Quantification of Ultrastructure from Transmission Electron Microscopy Images
Anqi Zou, Winston Tan, Jiayi Ji, Florencia Rojas-Miguez, Laura Dodd, Emily Oei, Sarah R. Vargas, Stephen P. Berasi, Hongying Yang, Hui Chen, Joel M. Henderson, Xueping Fan, Weining Lu, Chao Zhang
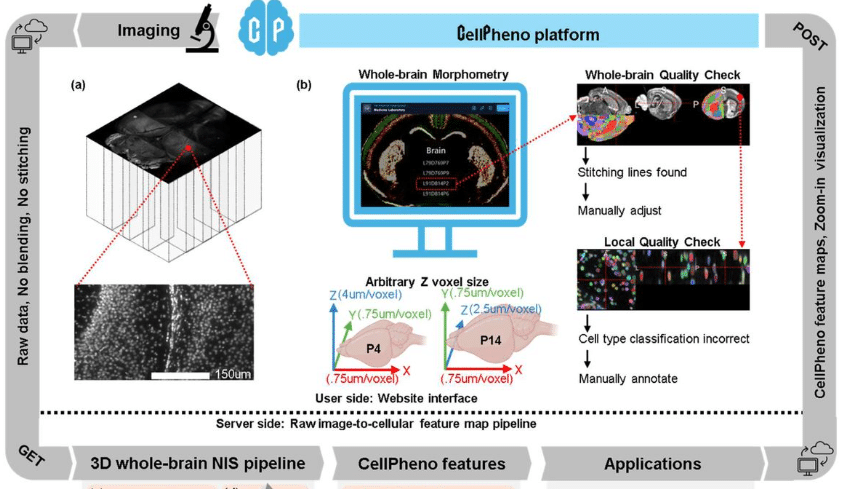
CellPheno: A High-throughput Computational Platform for Quantifying Cellular Resolution Whole Brain Microscopy Images
Ziquan Wei, Ian Curtin, Felix A Kyere, David Borland, Hong Yi, Minjeong Kim, Mustafa Dere, Carolyn M McCormick, Oleh Krupa, Yen-Yu Ian Shih, Mark J Zylka, Jason L Stein, Guorong Wu
NucleoNet and DropNet: Generalist deep learning models for instance segmentation of nuclei and lipid droplets from electron microscopy images
Abhishek Bhardwaj, Chris W. Dell, Melissa R. Mikolaj, Helen Spiers, Adam Harned, Balamurugan Kuppusamy, Peng Liu, Donglai Wei, Esta Sterneck, Kedar Narayan
micromorph: a Python toolkit for measurement of microbial morphology
Simone Coppola, Séamus Holden
SIMBA: an agentic AI platform for single-molecule multi-dimensional imaging
Hongjing Mao, Harsh Mauny, Obblivignes KanchanadeviVenkataraman, Caroline Laplante, Dongkuan (DK) Xu, Yang Zhang
Recovering membrane interaction kinetics of single molecules from 3D tracking data
Erik Lundin, Ivan L. Volkov, Magnus Johansson
PixelDeck: A local-first media library manager for biomedical imaging
Benjamin L Kidder
Sampling-Aware 3D Spatial Analysis in Multiplexed Imaging
Ido Harlev, Tamar Oukhanov, Raz Ben-Uri, Leeat Keren, Shai Bagon
MTCurv: Deep learning for direct microtubule curvature mapping in noisy fluorescence microscopy images
Achraf Ait Laydi, Sidi Mohamed Sid’El Moctar, Yousef El Mourabit, Hélène Bouvrais
Background Remover — an effective tool for processing noisy microscopy images
Anna Kilian, Paweł Bilski, Małgorzata Sankowska
Tailored Speckle Illumination Microscopy with Enhanced Sectioning and Image Quality
SeungYun Han, KyeoReh Lee, Young Seo Kim, Chuan Li, Nicholas Bender, Kabish Wisal, Taeyun Ku, Jerome Mertz, Hui Cao
ToFiE, a Topology-aware Fiber Extraction workflow for 3D reconstruction of dense and heterogeneous biological fiber networks from microscopy images
Risa Togo, Sara Cardona, Irène Nagle, Gijsje H. Koenderink, Behrooz Fereidoonnezhad, Mathias Peirlinck
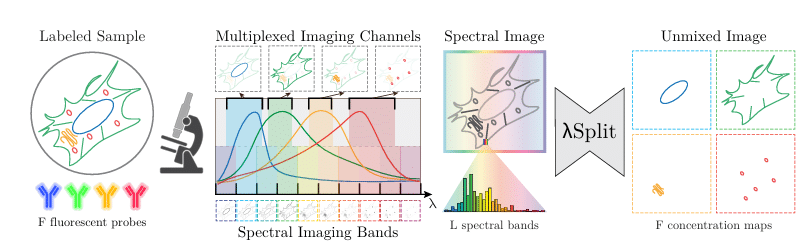
λSplit: Self-Supervised Content-Aware Spectral Unmixing for Fluorescence Microscopy
Federico Carrara, Talley Lambert, Mehdi Seifi, Florian Jug
Boundary-Aware Instance Segmentation in Microscopy Imaging
Thomas Mendelson, Joshua Francois, Galit Lahav, Tammy Riklin-Raviv


 (No Ratings Yet)
(No Ratings Yet)