Featured image with Virginia Barrera
Posted by FocalPlane, on 2 February 2024
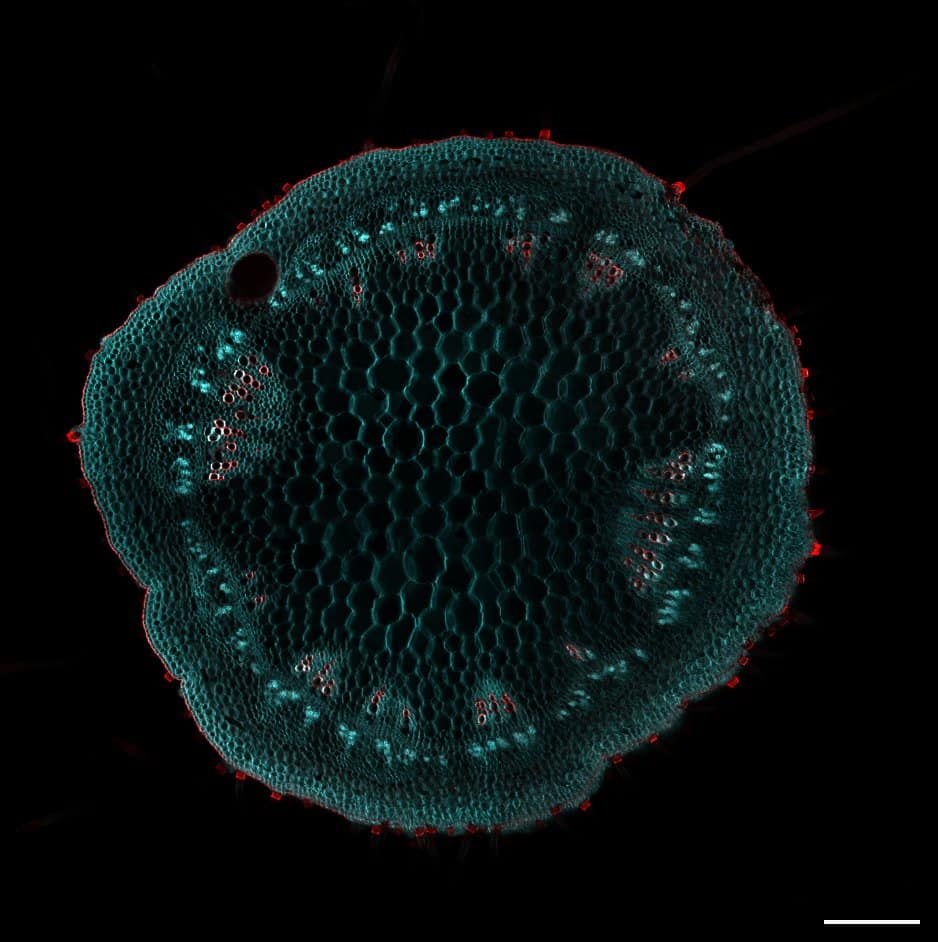
Our featured image was acquired by Virginia Barrera and was the winning image from the Institute of Molecular and Cellular Biology of Rosario (IBR-CONICET) annual retreat. IBR-CONICET was established in 1985 and is home to scientists researching different aspects of microbiology, biomedicine, structural biology, and agrobiotechnology. Virginia’s image depicts a transversal cut of a soybean, aiming to explore variations in cell architecture among different variants. To minimize the impact of autofluorescence in plants, they employed the ClearSee protocol for sample fixation, clarification, and staining. They used Calcofluor White to stain cellulose and Basic Fuchsine for lignin. This staining combination allowed them to distinguish cell sizes and shapes in the shoot, highlighting xylem and fiber structures. Following the preparation, Virginia mounted shoot dissections in ClearSee and visualized them using Laser Scanning Confocal Microscopy (LSCM).
Find out more about Virginia’s research below:
Research career so far: I earned my degree in Biotechnology from the National University of Rosario. Currently, I am pursuing a PhD at the Institute of Molecular and Cellular Biology of Rosario in Argentina, under the guidance of Dr Ramiro Rodriguez. Our group aims to analyze the cellular development and growth of plant organs to identify mechanisms regulating cell proliferation, expansion, and differentiation. Since the beginning of my PhD, I have extensively utilized microscopy to study the localization, duration, magnitude, and direction of these processes. While our primary focus is on the Arabidopsis model, we also explore new plant species and seek to translate our knowledge to agriculturally relevant species.
Current research: The final size of plant organs is determined by the number and size of their cells. Typically, cell number is controlled by the mitotic cell cycle, while cell expansion determines the final size. In my current research, we focus on unravelling the mechanisms that govern these processes, aiming to identify new regulators and understand the interactions between these pathways. To achieve this, we utilize time-lapse microscopy to analyze the progression of cell number and size in different plant organs during development and investigate how different genes can modify these processes.
Favourite imaging technique/microscope: Given the dynamic nature of the processes we study; I use time-lapse microscopy extensively. I particularly enjoy conducting experiments with this technique because it serves as a reminder that the images captured represent specific moments, and a lot of events leading up to that point and will unfold afterward. It adds a fascinating dimension to the understanding of the biological processes under observation.
What are you most excited about in microscopy? I’m particularly enthusiastic about microscopy techniques that enable the tracking of living cells. Currently, I’m doing an internship at the Routier Lab at Montreal University, where I’m training in MorphoGraphX, a 3D image analysis software enabling the quantification of growth, cell shape, gene expression, and mechanics. I believe that the concurrent development of tools facilitating 3D visualizations and analysis of large image datasets is leading to numerous and profound advancements in the field.


 (No Ratings Yet)
(No Ratings Yet)