A new version of TrackMate with improved integration of key AI algorithms and track edition improvements
Posted by Jean-Yves Tinevez, on 12 December 2025
We’re excited to announce TrackMate v8.1, the latest version of Fiji’s powerful cell and organelle tracking plugin. This release combines all the major features from v8 with important usability improvements, making TrackMate more useful and easier to use than ever before.
What is TrackMate?
TrackMate is an extensible, open-source tracking software designed for biologists and microscopy researchers to analyze moving objects in time-lapse images. As a plugin for Fiji, it provides an accessible yet sophisticated solution for tracking cells, organelles, particles, and other biological structures in both 2D and 3D microscopy data.
With TrackMate, you can:
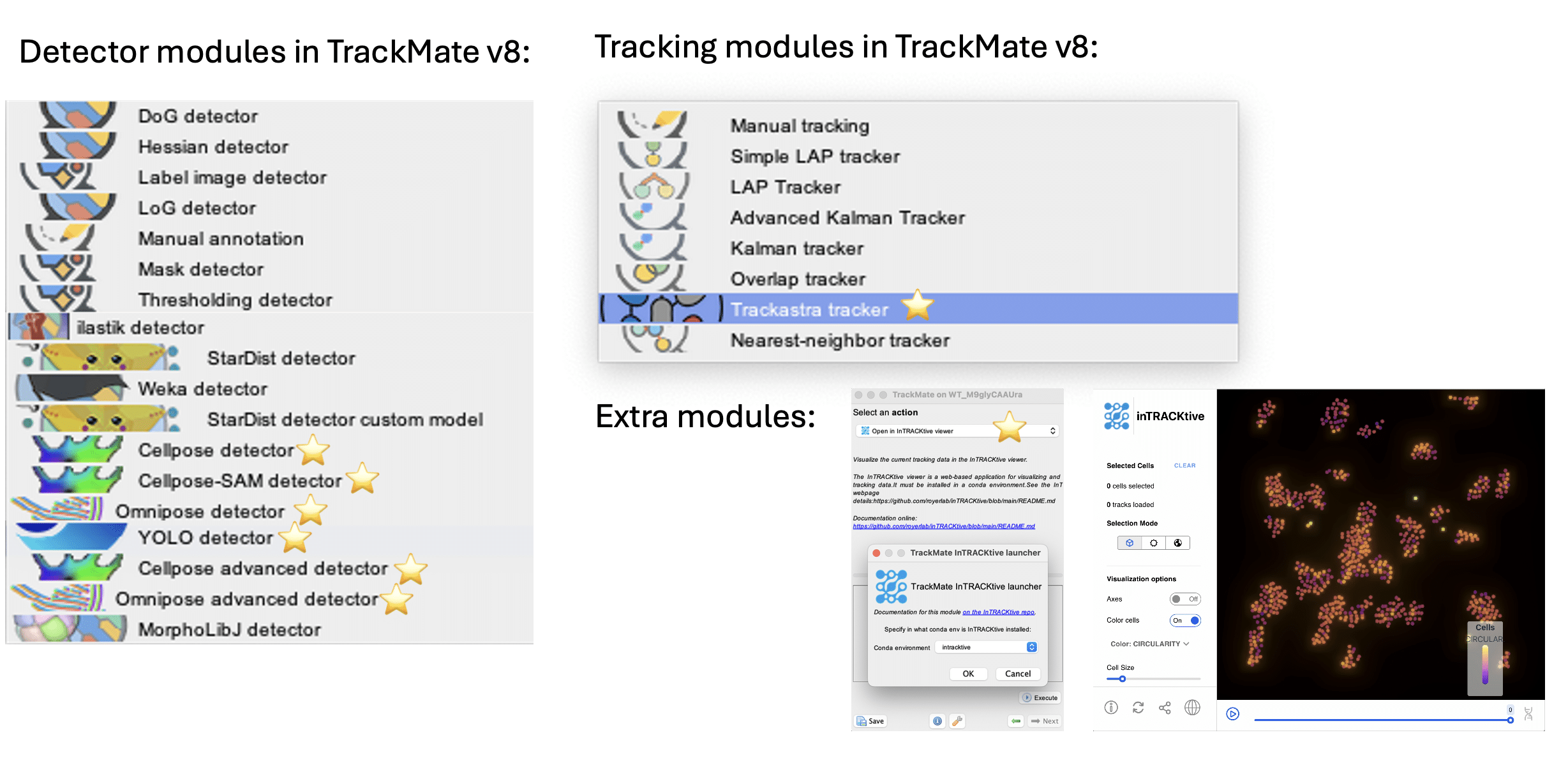
- Detect objects automatically using state-of-the-art AI algorithms (Cellpose, Omnipose, YOLO, StarDist, etc.), classical algorithms (LoG, mask, label images)
- Track movements over time with advanced linking algorithms (LAP tracker, Kalman tracker, Trackastra, etc.)
- Manually edit or curate tracking results. TrackMate lets you combine automatic, semi-automatic or fully manual approaches for your data
- Analyze trajectories to measure velocity, displacement, morphology changes, and other quantitative features
- Edit results with intuitive tools for manual correction and ground-truth generation
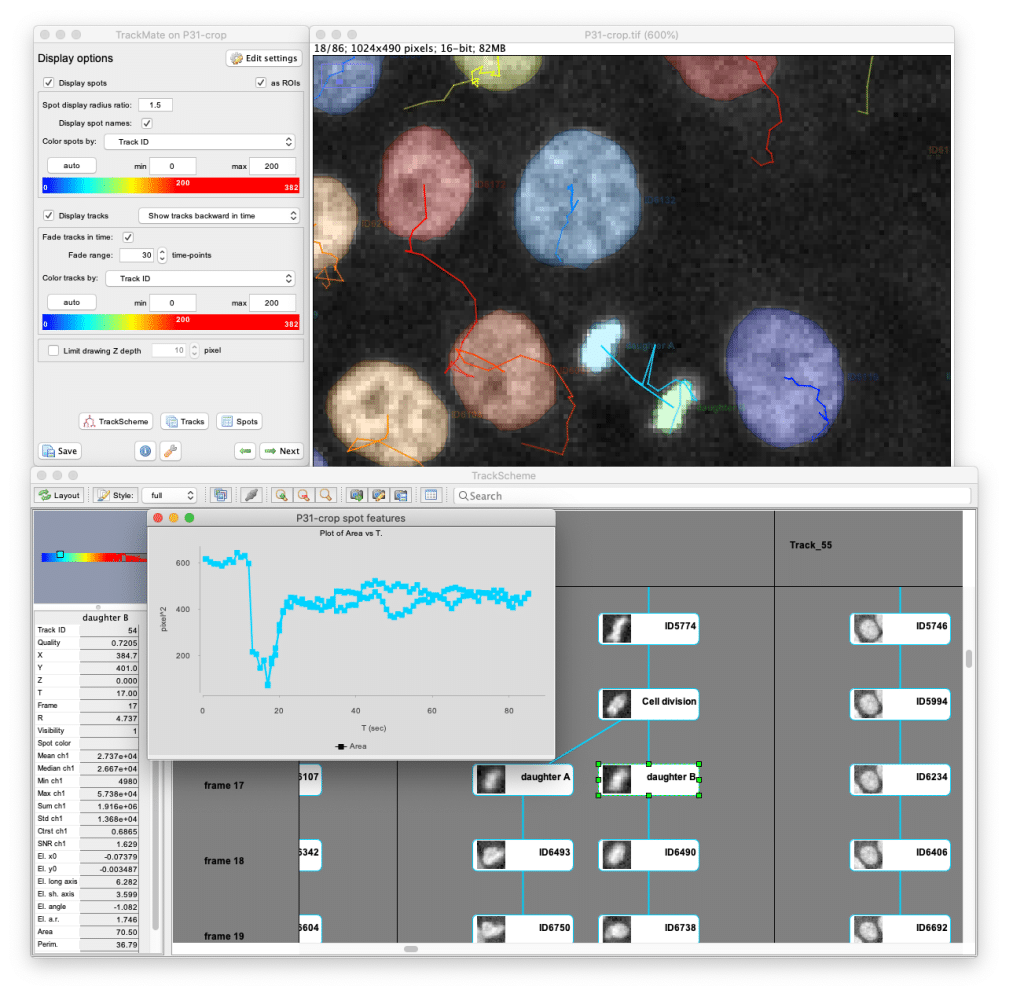
- Visualize data through multiple interfaces (TrackScheme for lineage analysis, etc.)
- Export results for further analysis in Python, MATLAB, or other software
TrackMate handles both simple spot-like objects (vesicles, nuclei) and complex cell-like objects with detailed shape information. Its modular design allows users to choose from multiple detection and tracking algorithms, while developers can easily integrate new methods. The software includes powerful analysis tools to compute numerical features, generate plots, and create publication-quality visualizations of your tracking data.
Whether you’re studying cell migration, organelle dynamics, bacterial movement, or developmental processes, TrackMate provides a complete pipeline from raw images to quantitative results – all within the familiar Fiji environment.
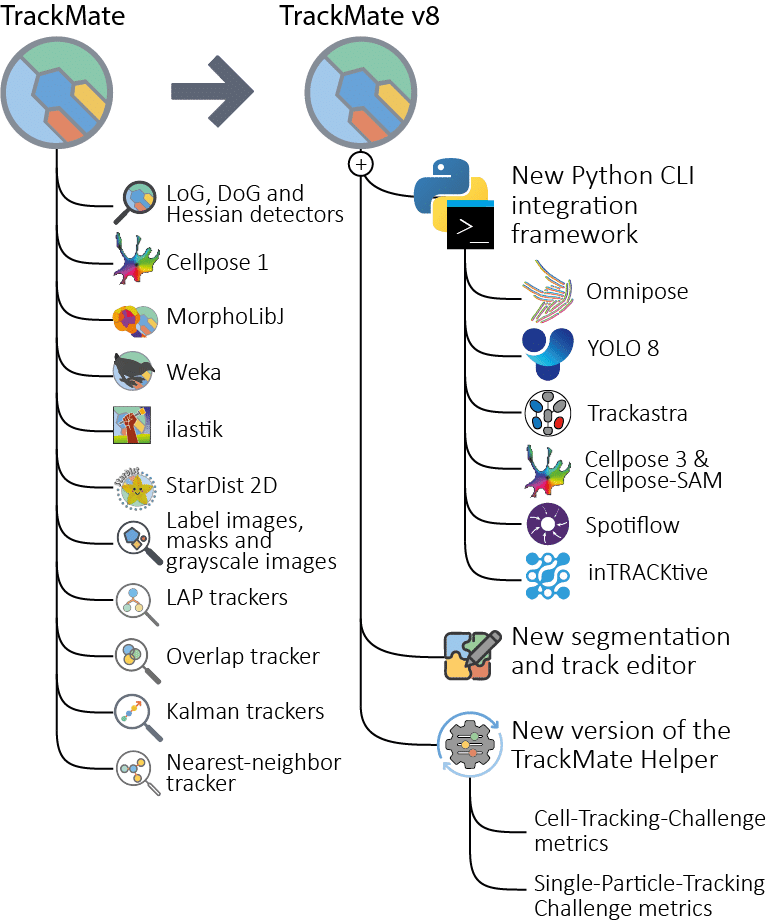
Key new features in the v8 version
2D Segmentation Editor with Advanced Editing Tools
TrackMate now includes a comprehensive 2D segmentation editor based on LabKit, allowing you to create and manually edit object shapes in 2D by painting them over the input image. Combined with the link editing capabilities of TrackScheme, this feature enables TrackMate users to:
- Generate ground-truth tracking data from scratch
- Refine automated segmentation results.
It can handle overlapping objects seamlessly, and several editing modes have been introduced to facilitate and accelerate editing of object shapes.
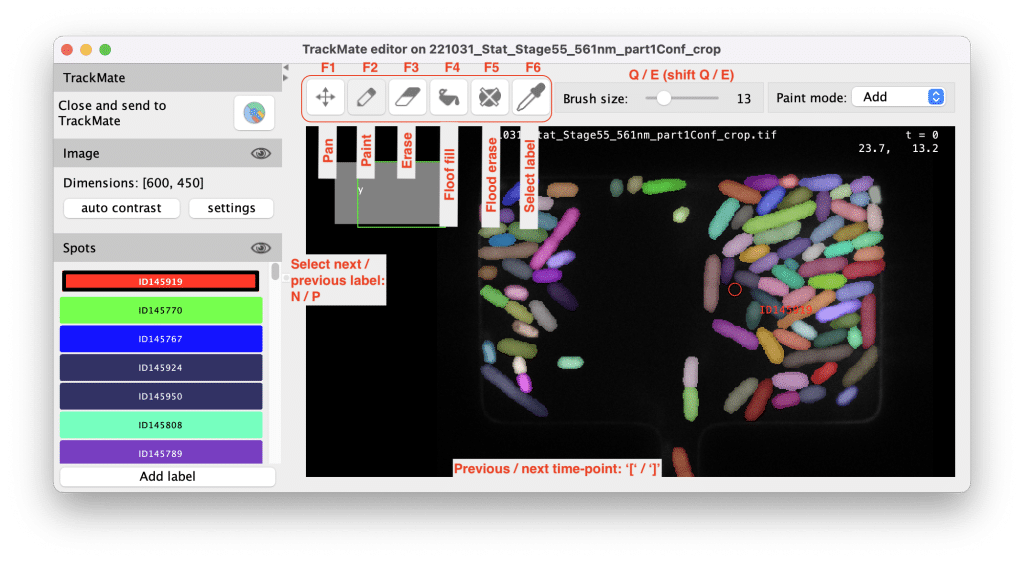
This editor is based on Labkit components and works great e.g. with a tablet. For convenience, all key bindings can be redefined on the fly. Documentation and examples can be found here.
Better Python integration support
TrackMate v8 features a completely reworked framework for integrating Python-based detection and tracking algorithms, with support for:
| Algorithm | Type | Key Features |
|---|---|---|
| Cellpose | Segmenter | Includes Cellpose-SAM, and support for Cellpose v3. |
| Omnipose | Segmenter | Optimized for bacteria / filaments |
| YOLO | Detector | A popular AI-based object detection as spot detector |
| Trackastra | Tracker | Advanced AI-based tracking algorithm |
| Spotiflow | Detector | Robust AI-based spot detection |
| inTRACKtive | Tracker | In browser tracking data visualization. |
These integrations are based on a new development framework to integrate with Python tools installed and managed within conda environments. It will make running these algorithms more robust on all major OSes. TrackMate still needs to know where conda is installed, and a new configuration tool automatically detects conda installations, handling a large number of edge cases.
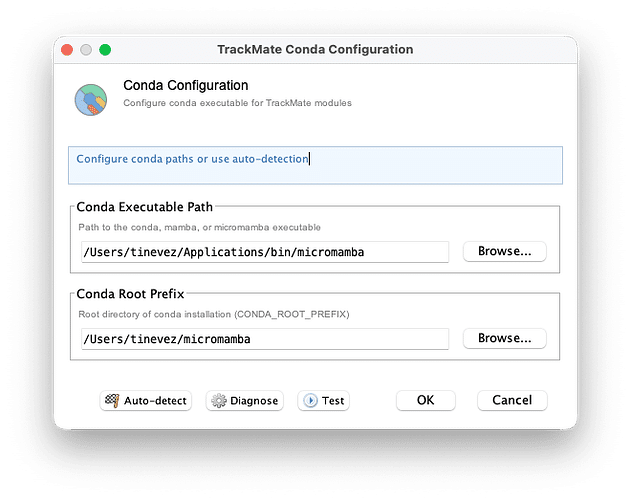
For developers:
This new framework will considerably accelerate the integration of detection and tracking algorithms in TrackMate. It automates the parameters I/O, the creation of the UI and all the boilerplate code. For instance, integrating Spotiflow took us less than an afternoon.
The framework is located in the fiji.plugin.trackmate.util.cli package. Developers can extend the CLIConfigurator class to define CLI parameters, using a syntax similar to Python’s argparse.
Example: Trackastra Integration
public class TrackastraCLI extends CondaCLIConfigurator {
// ...
public TrackastraCLI(final int nChannels) {
this.modelPretrained = addChoiceArgument()
.name("Model pretrained")
.help("Name of pretrained Trackastra model.")
.argument("--model-pretrained")
.addChoice("general_2d")
.addChoice("ctc")
.defaultValue(DEFAULT_TRACKASTRA_MODEL)
.key(KEY_TRACKASTRA_MODEL)
.get();
this.customModelPath = addPathArgument()
.name("Path to a custom model")
.argument("--model-custom")
.help("Local folder with custom model.")
.defaultValue(DEFAULT_TRACKASTRA_CUSTOM_MODEL_FOLDER)
.key(KEY_TRACKASTRA_CUSTOM_MODEL_FOLDER)
.get();
// ...
}
}
GEFF early support
GEFF is a new, universal file format for exchanging tracking data between tools. While Java support is still in beta, it’s already used in TrackMate (e.g., for the TrackMate-inTRACKtive integration). This was done with Laura Xénard, Ko Sugawara, Tobias Pietzsch and all the great minds behind LiveImageTrackingTools.
Better tools for tracking parameter optimization
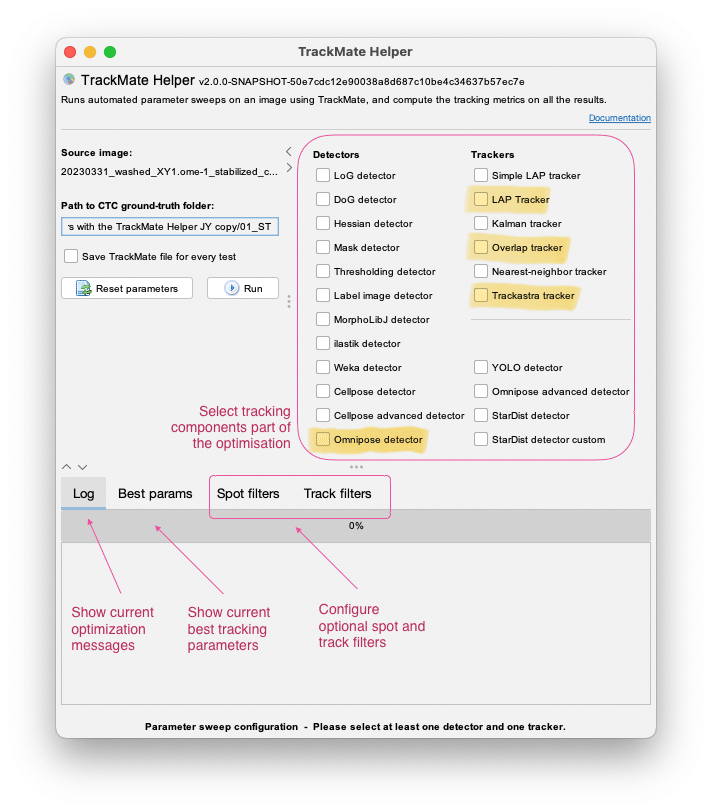
The TrackMate-Helper tool is a tool that can sweep over an extensive range of user-defined tracking parameters, including detectors and tracking algorithms, to define the best possible combination of tracking parameters for your tracking task. This might become important, because now you have a large number of algorithm combinations and parameterizations to choose from:
With this release, we rewrote the TrackMate-Helper in full, adding:
- New tracking metrics for the optimization:
- Single Particle Tracking (SPT) Challenge metrics
- Cell Tracking Challenge (CTC) metrics
- The inclusion of spot and track filters in the parameter sweeps
- Several fixes improving performance and robustness
Its adjunct tool the TrackMate-Batcher offers a user interface to run tracking on a batch of images. It has been rewritten with a few minor quality of life features.
Reworked the Cellpose and new Omnipose integration
Using the new Python integration framework mentioned above, we revisited its integration in TrackMate. Now the ‘TrackMate-Cellpose’ module offers:
- Cellpose 3: Easier installation, GPU support everywhere.
- Cellpose-SAM: the version 4 of Cellpose based on SAM. This new version of Cellpose requires much less parameter tuning. I have included one: the possibility to pass to the model one channel or all channels. This might impact the segmentation accuracy based on your input, as illustrated here.
- And Omnipose: For bacterial and filamentous cells
Documentation: Omnipose Cellpose Cellpose-SAM
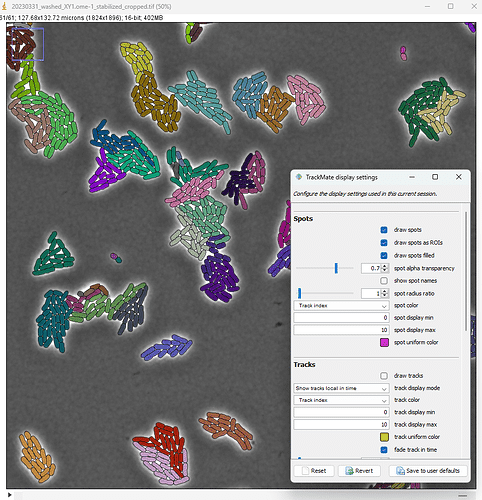
Trackastra integration
Trackastra can be used as a tracker in TrackMate!
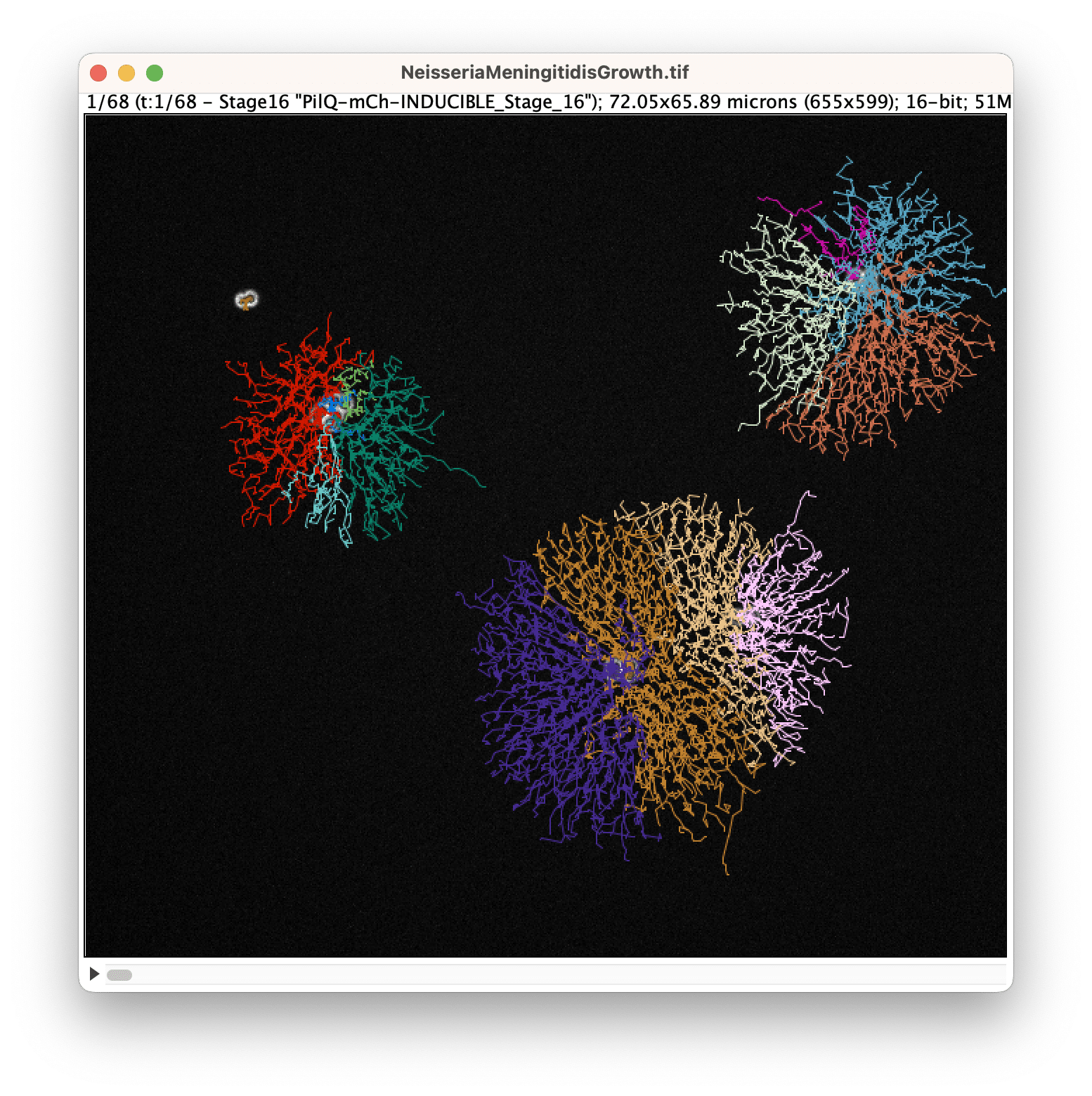
YOLO integration
We have added the popular YOLO detection algorithm as a spot detector in TrackMate. Here we integrated the object detection routine of YOLO, so objects are detected as boxes and add in TrackMate as circular spots. YOLO does not ship pretrained model for microscopy, so you will have to provide or train one yourself. We have made an example use of YOLO to track bacteria against a complex background, to be released soonish.
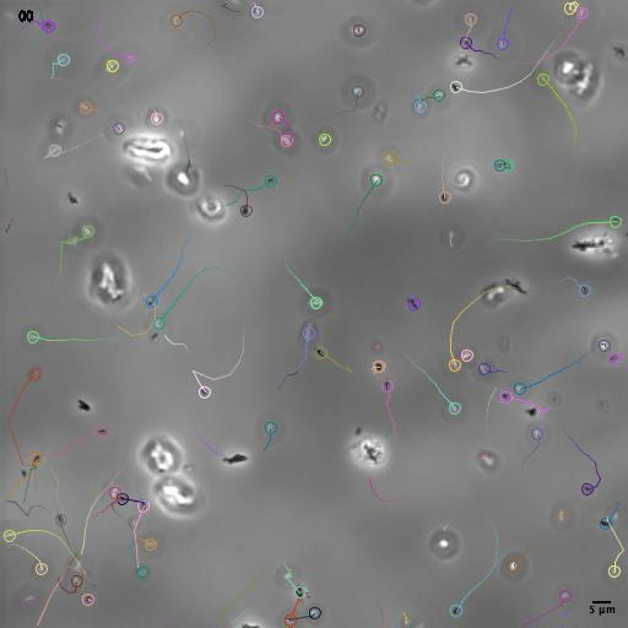
Spotiflow
Spotiflow is an AI based spot detection algorithm, made to be robust against imaging defects. Its integration in TrackMate allows for using it seamlessly in tracking projects.
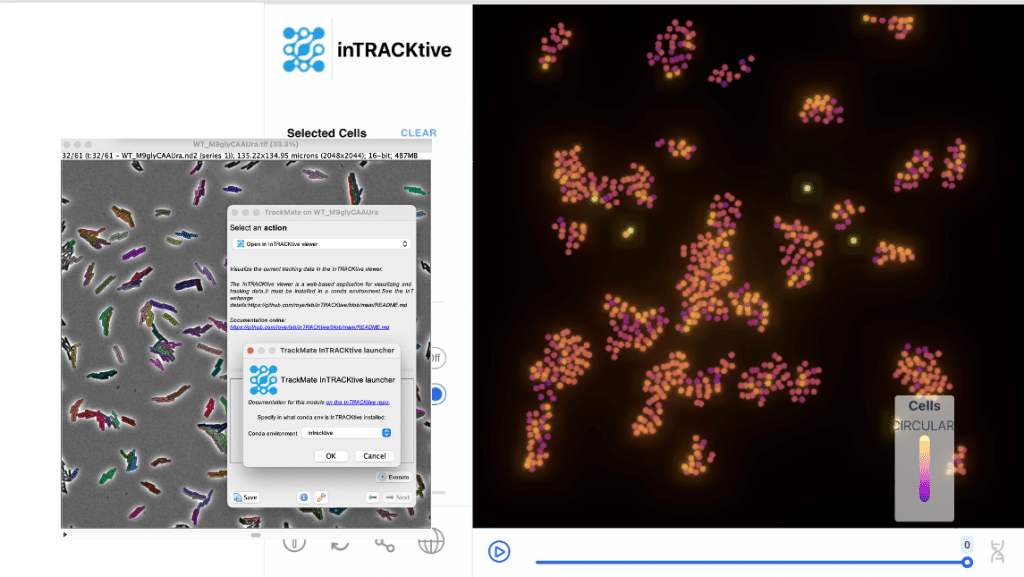
inTRACKtive
You can now use inTRACKtive as an external viewer for your tracking data. InTRACKtive is a web-based browser made for large tracking data exploration.
Additional improvements
Some changes are less visible but equally important. In particular, we have fixed a long-standing memory leak issue, which required major changes in the API. If you are a developer and maintain a TrackMate detector or segmenter module, there is a good chance that the v8 will break it.
Acknowledgments
TrackMate enjoys a lively and friendly community. For this release we have been lucky to collaborate with:
- For Omnipose: Kevin Cutler, Marie Anselmet, Gaëlle Letort
- Trackastra: Benjamin Gallusser and Martin Weigert
- inTRACKtive: Teun Huijben
- Spotiflow: Albert Dominguez
- GEFF: Tobias Pietzsch, Laura Xénard, Ko Sugawara
- And the F-BIAS team for beta testing!
TrackMate has been in development within the Institut Pasteur Paris since a decade or so, collaborating with excellent people everywhere. In particular
- Curtis Rueden and Kevin Eliceiri in the University of Madison, who also supports the whole Fiji ecosystem.
- Guillaume Jacquemet’s team in the University of Turku.
Get started
- Download a fresh ‘Fiji-latest‘ at https://fiji.sc. This is important as the new version if not shipped in the previous version of Fiji based on Java 8.
- Read the documentation
- Share your feedback and use cases on the Image-SC forum!
We’re excited to see how these new features will enhance your tracking workflows!


 (5 votes, average: 1.00 out of 1)
(5 votes, average: 1.00 out of 1)