Imaging spotlight: PhotoFiTT
Posted by FocalPlane, on 2 March 2026
In this paper highlight, we learn about PhotoFiTT, a framework for measuring phototoxicity in live-cell fluorescence microscopy. Mario Del Rosario and Estibaliz Gómez-de-Mariscal, the co-first authors of PhotoFiTT answer our questions about their work.
What is PhotoFiTT?
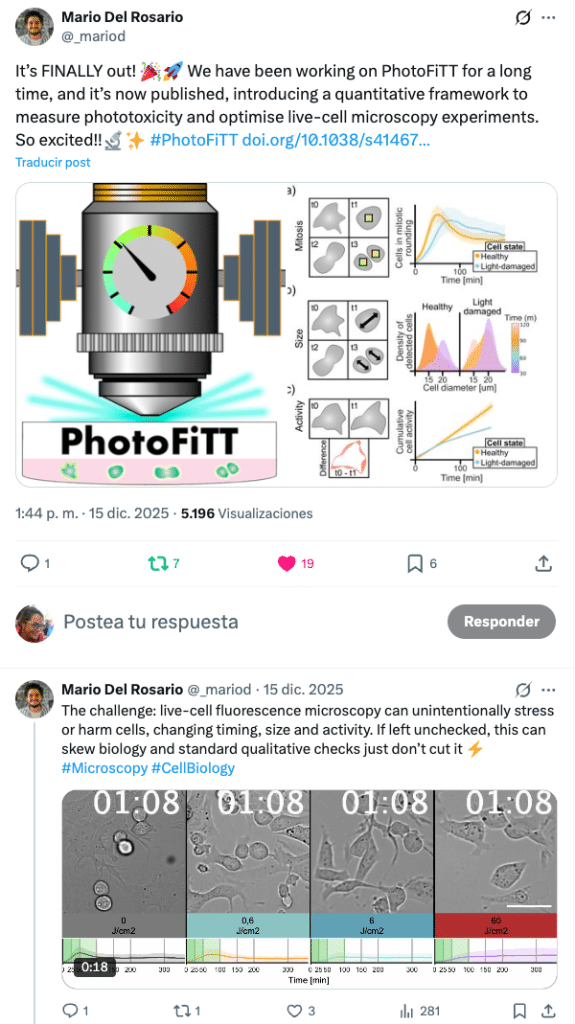
PhotoFiTT (Phototoxicity Fitness Time Trial) couples a standardised live‑cell label-free imaging protocol with AI analysis to quantify the effects of phototoxicity in a highly generalisable manner. Using deep learning, it measures deviations in the physiological cell-clock (cell cycle) and uses them as quantitative readouts of photodamage. This way, PhotoFiTT transforms the question “am I frying my cells?” into a dose–response experiment, allowing for the optimisation of live-cell microscopy acquisitions, reporting of phototoxicity and improvement of experimental reproducibility. We applied it to different wavelengths, light doses and imaging approaches, and observed shifts in division timing and cellular activity. To support adoption, PhotoFiTT is released as a user-friendly, open-source workflow on GitHub, allowing researchers to analyse similar time-lapse datasets beyond those presented in the manuscript.
What made you embark this project?
The project started from a very practical frustration. Phototoxicity is something all live-cell microscopists worry at some point in their experiments but the tools to assess it are either too subjective or indirect. People often rely on visual inspections, as in, “the cells look fine, let’s carry on” or on stress markers that themselves require extra labelling and illumination.
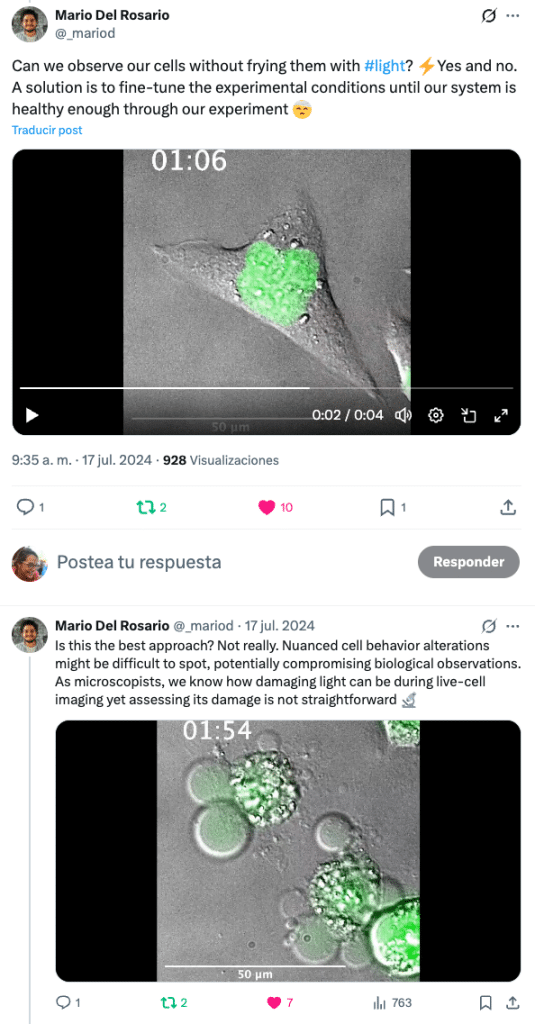
We wanted a way to quantify phototoxicity that was biologically meaningful, label-free and compatible with how live-cell microscopy is actually done. Cell cycle timing emerged as a powerful and underused readout: cells integrate stress over time, and even subtle photodamage shows up as delays or altered dynamics long before overt cell death.
We also wanted a highly reproducible and relatively easy to implement setup. Current AI techniques and user-friendly tools to train models (e.g., DL4MicEverywhere) support the quick adoption and automatic image processing, fostering a highly quantitative and reproducible pipeline. PhotoFiTT grew from that idea.
How it has been used so far?
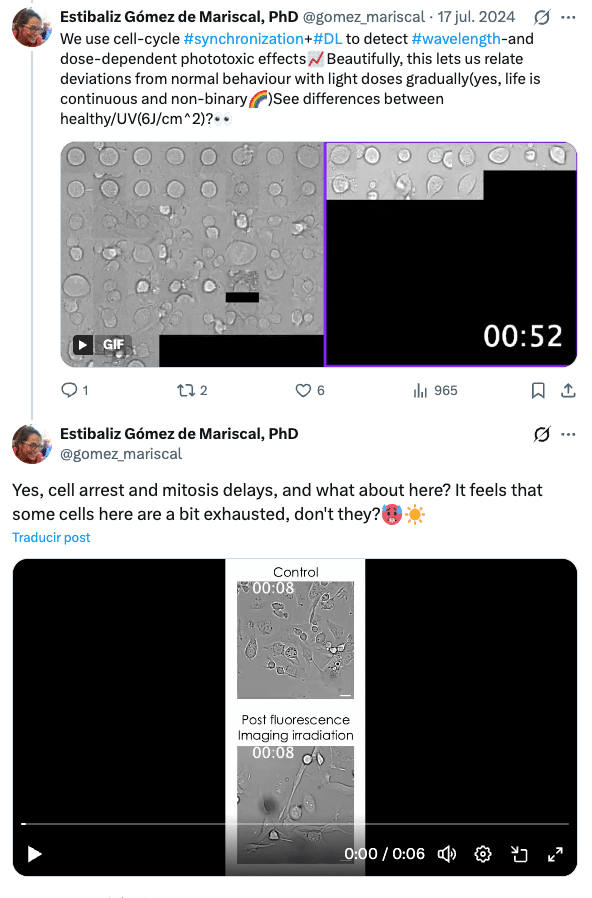
We assess the effects of phototoxicity in adherent mammalian cells (CHO and HeLa) to map phototoxicity across near‑UV (385 nm), green (475 nm, GFP) and red (688 nm) excitations, showing, for example, that 385 nm can already cause strong mitotic delays at doses around 0.6 J/cm² (used in lattice light‑sheet and widefield microscopy modalities), while longer wavelengths require higher doses to reach similar impact. The framework quantified continuous changes in mitotic delay, daughter cell appearance and cumulative activity across a range of exposures, demonstrating that phototoxicity is graded rather than purely lethal vs non‑lethal. We also inspected the increasing photodamaging effect of illuminating cells that have already been labelled (e.g., MitoTracker) or with different imaging modalities (e.g., widefield, confocal spinning disc). These experiments show how PhotoFiTT can benchmark different fluorescence microscopy imaging conditions (wavelength, light dose and exposure frequency) and directly link them to cell cycle and behavioural outcomes.

Who else it could be useful for?
Anyone studying cell behaviour in long‑term or high‑intensity live‑cell imaging setups (e.g., cell migration, mitosis, morphogenesis, or drug‑response assays) can benefit from using PhotoFiTT. Microscopists developing or optimising new imaging modalities (e.g. lattice light‑sheet, high‑NA spinning disk, light‑sheet variants) can also use it as a quantitative “stress test” for illumination strategies. Moreover, novel smart microscopy approaches aiming to push some of the limitations of live-cell microscopy, can also use this as a baseline to implement more efficient imaging approaches.
Moving beyond, we believe that PhotoFiTT is also relevant for core facilities and imaging standards initiatives aiming to report phototoxicity in a comparable manner. For reviewers or method developers who need robust evidence that imaging protocols preserve cell fitness or that the experimental outcomes are not compromised by phototoxicity. More broadly, any lab that cares about reproducibility in live‑cell microscopy can integrate PhotoFiTT to define and document safe illumination windows for their favourite cell lines and fluorophores.
What about experiments on non-adherent cells?
Adherent mammalian cells were a natural starting point because mitotic timing is easy to measure and highly informative. However, the core PhotoFiTT’s logic of using intrinsic biological timing or behaviour as a fitness readout is not limited to these systems.
With appropriate adaptations, the framework could be extended to other cell types or biological contexts, as long as a reproducible, label-free biological event can be detected. For example, this could be integrating morphodynamic metrics, or other label‑free markers of stress that are compatible with the non‑invasive philosophy. Exploring these extensions is an important direction for future work.

What do researchers need to be able to use PhotoFiTT?
Experimentally, researchers need a live‑cell imaging setup with a cell line that can be imaged in transmission (for label‑free phase‑contrast) during and after the photodamage pulse. In our protocol, cells are synchronised at G2/M (using a CDK1 inhibitor) to make mitotic timing a sensitive readout, but PhotoFiTT’s logic and analytics are applicable to other systems as long as division events and timing are measurable.
Computationally, a basic Python or Conda installation and the user-guide in PhotoFiTT’s GitHub repository is sufficient to implement PhotoFiTT. The analysis and data visualisation are fully reproducible following the steps in PhotoFiTT’s notebooks.

What different techniques did you bring together when working on PhotoFiTT?
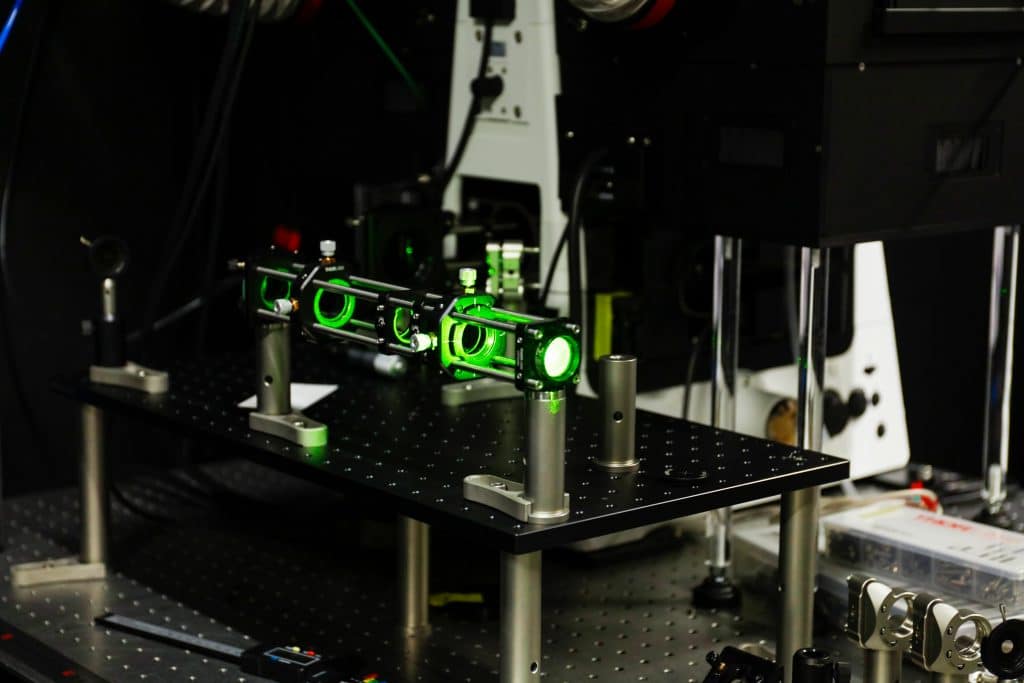
PhotoFiTT is very much a hybrid project. Experimentally, it relies on careful live-cell imaging and controlled illumination. We used 3D printed components that helped us accurately reproduce the experimental pipeline across lab members. On the analysis side, AI enables robust detection of cells and mitotic events without fluorescent labels.
This combination allows us to connect physical imaging parameters such as wavelength, dose and modality with biological consequences in a quantitative way. Importantly, none of the individual techniques are exotic on their own; it’s their integration into a coherent workflow that makes the framework broadly applicable.

What are the challenges of phototoxicity?
Understanding phototoxicity sits at the intersection of physics, biology and experimental practice. One of the biggest challenges though, is that it is often invisible until it is too late. Cells can look morphologically normal while already being stressed, delayed, or functionally altered. This means that biological conclusions can be subtly biased without researchers realising it. As imaging technologies become faster, brighter and more complex, the risk of unintentionally perturbing the system increases. Quantifying phototoxicity is therefore essential for both experimental optimisation and scientific reproducibility.
Where can readers find more information?
The full peer‑reviewed manuscript is available in Nature Communications under the title “PhotoFiTT: A quantitative framework for assessing phototoxicity in live‑cell fluorescence microscopy”, containing all the information about PhotoFiTT. Additionally, we have a GitHub page that can provide everything that’s needed to start using the tool.


 (No Ratings Yet)
(No Ratings Yet)