Imaging spotlight: PLIN5 phosphorylation tunes mitochondria–lipid droplet coupling in the liver
Posted by FocalPlane, on 29 April 2026
In this paper highlight, we hear from Natalie Porat-Shliom, who discusses her recent paper on how PLIN5 phosphorylation regulates mitochondria–lipid droplet contacts in the liver.
What are the key results from your paper?
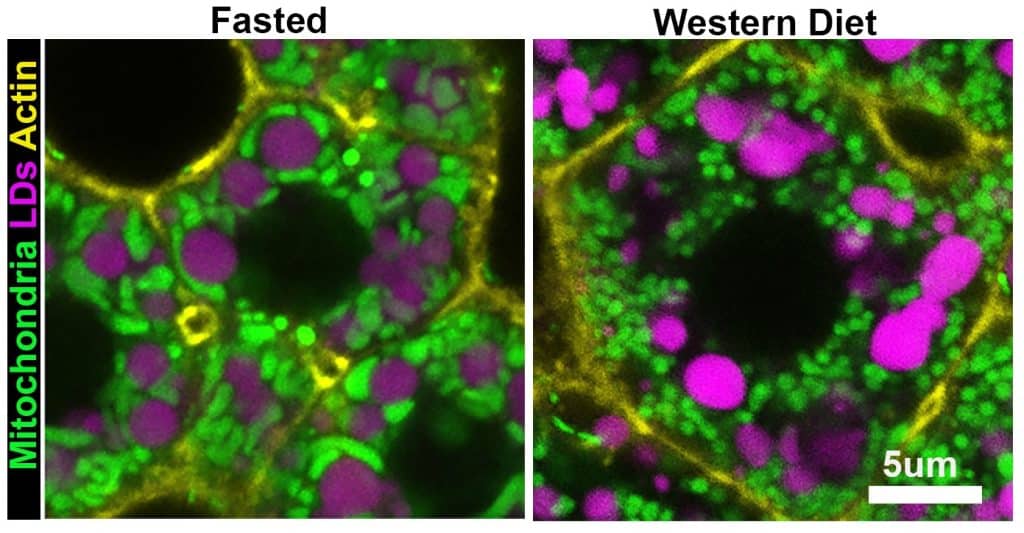
Our study asked how hepatocytes handle a sudden increase in lipid flux. We found that fasting and a Western diet both raise liver lipid levels, but they do not trigger the same organelle response. In fasted mice, mitochondria and lipid droplets were extensively remodeled across the lobule, and mitochondria–lipid droplet contacts increased markedly. In contrast, in mice fed a short-term Western diet, those contacts were relatively infrequent, even though the liver was also becoming steatotic. This told us that not all lipid-rich states are the same at the organelle level.
To understand what controls this response, we combined imaging with spatial proteomics and identified PLIN5 as a strong candidate. PLIN5 was upregulated in the fasted liver and localized to the mitochondria–lipid droplet interface. We then showed that its phosphorylation state acts like a switch: the phospho-null mutant PLIN5-S155A promoted mitochondria–lipid droplet contacts and enlarged lipid droplets, whereas the phospho-mimetic PLIN5-S155E reduced contacts and produced fewer, smaller droplets. PLIN5-S155A also promoted triglyceride storage and reduced lipotoxicity and oxidative stress during short-term lipid overload.
With prolonged Western diet exposure, and in human liver samples with higher lipid content, mitochondria–lipid droplet interactions became more abundant, pointing to a dynamic role for these structures in liver disease.
Which imaging and image analysis techniques have you used in your research? Can you tell us more about scPhenomics?
We used confocal tile scans and z-stacks of liver sections from mtDendra2 mice, with fluorescently labelled mitochondria. Lipid droplets were stained with LipidTox and cell structure with phalloidin. Imaging was performed on a Leica SP8 confocal microscope, and we used high-resolution z-stacks and 3D surface rendering in Imaris to validate our measurements and analyze human liver samples.
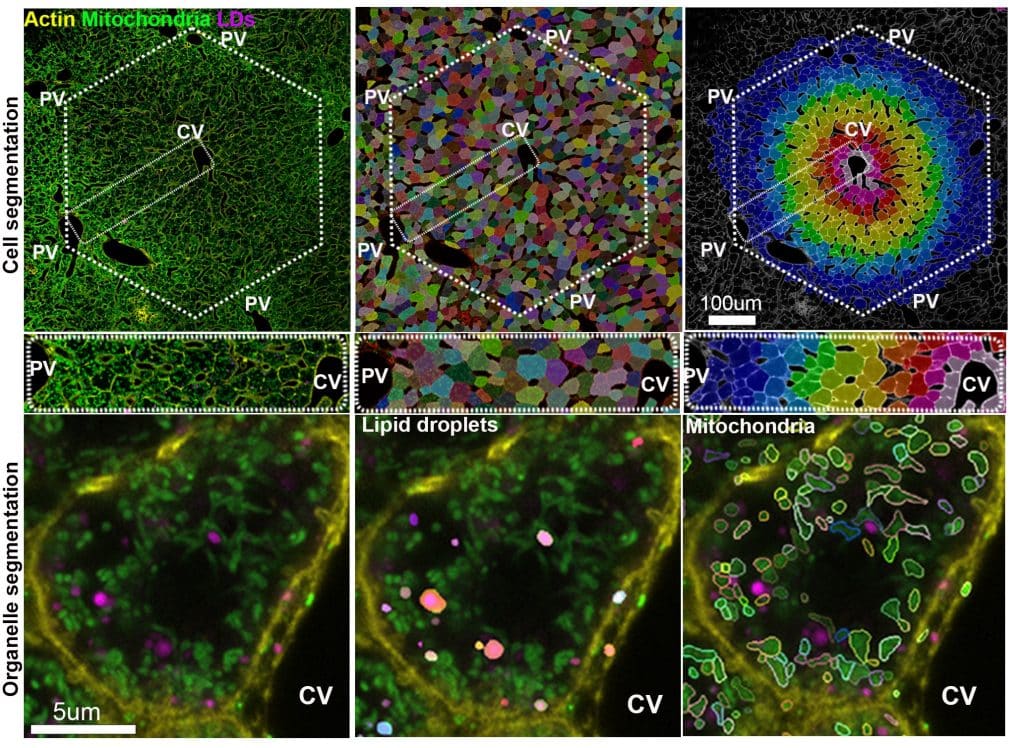
scPhenomics is the analysis framework we developed to extract single-cell phenotypes while preserving tissue context. What makes scPhenomics especially powerful is that it captures how neighboring hepatocytes can look and behave very differently. Rather than averaging the liver response using traditional biochemical methods, scPhenomics lets us map each hepatocyte along the periportal-to-pericentral axis and quantify how organelle features vary across space and metabolic states. The pipeline uses a custom Python workflow with Cellpose models to segment hepatocytes, mitochondria, and lipid droplets and then measure features such as size, shape, distance, and overlap.
Were there any technical challenges that you had to overcome when doing this research?
The biggest technical challenge was imaging across scales. We needed high enough resolution to capture mitochondria–lipid droplet interactions at the subcellular level, while also imaging large enough tissue areas to preserve spatial information across the lobule. This is challenging because high-resolution imaging takes time, especially when multiple channels are required, and even then, 2D images can create a false impression of organelle interactions, as these structures are small, complex, and three-dimensional.
We addressed this by combining throughput with validation. We used scalable 2D imaging for large-area analysis, but validated the key phenotypes using high-resolution confocal z-stacks and 3D Imaris reconstructions. Once we were confident in the phenotype, we focused on increasing the number of lobules, mice, and dietary conditions.
Are there any advances in imaging or image analysis that would help your research going forward?
Methods that combine large-volume 3D imaging with higher spatial resolution would be especially valuable. In this study, we relied on scalable 2D phenotyping supported by selected 3D validation datasets. Techniques that let us directly resolve contact-site architecture across larger tissue volumes would help us move from measuring proximity and overlap to measuring these interfaces more directly.
We would also benefit from approaches that connect morphology and molecular state in the same intact tissue. Future methods that integrate multiplexed imaging, spatial proteomics, and single-cell image analysis in situ would allow us to directly link organelle organization to cell state.


 (No Ratings Yet)
(No Ratings Yet)