Imaging spotlight: dissecting the precancerous niche
Posted by FocalPlane, on 25 March 2026
In this paper highlight we hear from Greta Skrupskelyte and Maria Alcolea, who discuss their recent paper dissecting the role of the microenvironment and tumour-stromal communication in tumour growth and survival.
What are the key results from your paper?
Research over the last decade has shown that the same mutations known to be linked to cancer, also accumulate in the tissues of healthy people during ageing. This has revealed an apparent contradiction; how can tissues harbour cancer-linked mutations and not develop cancer?
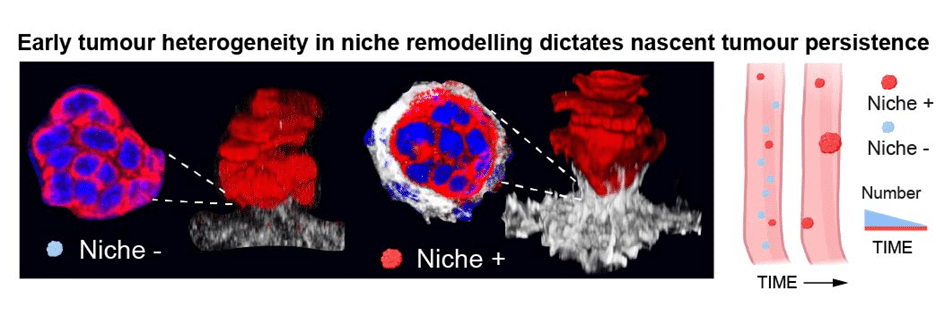
In our bodies most emerging tumours are eliminated, but some escape this elimination process. We asked how do tumours escape the clearance? Our study shows that the earliest epithelia–stroma interactions determine whether emerging tumours are cleared or allowed to persist. Surviving early tumours activate a stress response that mobilises fibroblasts. These cells remodel the extracellular matrix, building a fibronectin-rich “pre-cancerous niche”, a wound-healing programme gone rogue, that supports tumour growth and survival. Remarkably, this niche alone can induce tumour-like properties in otherwise normal epithelial cells. When we blocked tumour–stromal communication, far fewer tumours persisted. Surviving features are also found in human tumours opening new avenues for early detection and prevention.
Which imaging and image analysis techniques have you used in your research?
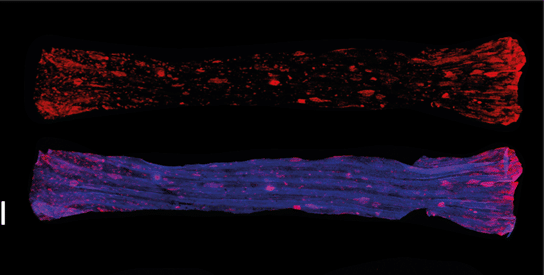
To study tumour emergence we have used cigarette smoke carcinogen and mouse oesophageal tissue. The oesophagus is a fortunate organ as the entire mouse oesophagus can be mounted on a single slide for imaging. Our unique sample processing and mounting techniques made the tumour quantifications across the entire tissue possible. Sample processing was an important aspect of successful imaging. The outer muscle from oesophagus was removed making the tissue around 200-250 um thickness. Tissue was mounted using 0.05mm spacers in RapiClear medium allowing tissue clearance. Lastly, tissue was mounted between two coverslips to allow imaging from both sides. Up to 4 replicate oesophagi were fitted in one imaging ‘slide’ (2 coverslips). The whole tissue scanning was performed using confocal microscopy (Stellaris 8); using the navigator mode tissues were mapped on a slide and an overnight ‘mark and find’ (each oesophagi was a separate object) was set up. Stellaris was our preferred platform as it gives great resolution, is very fast, and produces tile scans that are reasonably sized allowing easy manipulation/analysis of the raw files produced. For image analysis we have used Volocity and Arivis platforms, each with their unique advantages. Volocity allows quick overview of xyz in a single window while strolling through z plains, crucial for assessing incipient tumour niche (fibroblast involvement). It also allows image cropping in any direction (not just vertically or horizontally), enabling us to prepare images like the one attached. Arivis is incredibly good for a merged tile analysis, it often provided an overview map of the tissue, making sure no tumour is counted twice if analysis is done in individual tiles.
Are there any advances in imaging or image analysis that would help your research going forward?
Our published and many ongoing projects in the laboratory would greatly benefit from spatial proteomics. However, the platform is optimised for thin sections, while we work with wholemounts and perform whole tissue scans to identify the small clusters of cells for analysis. These clusters are not visible by the naked eye and, therefore, we cannot successfully section these unless it is done ‘blindly’. It would be an immense advance for our research to be able to use tissue whole mounts in multiplex immunofluorescence.


 (1 votes, average: 1.00 out of 1)
(1 votes, average: 1.00 out of 1)